Altered circadian expression of cytokines and cytolytic factors in splenic natural killer cells of Per1(-/-) mutant mice
- PMID: 23402528
- PMCID: PMC3595954
- DOI: 10.1089/jir.2012.0092
Altered circadian expression of cytokines and cytolytic factors in splenic natural killer cells of Per1(-/-) mutant mice
Abstract
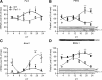
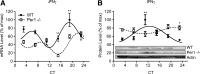
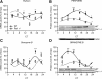
Circadian systems regulate the immune system by various molecular and physiological pathways. Disruption to the circadian temporality of these pathways is associated with disease formation and progression. Circadian clock genes have been shown to regulate pathways involved in cellular proliferation, apoptosis, and DNA damage response, as aberrant rhythms in these genes are associated with various diseases. However, there is growing evidence that specific circadian genes differentially regulate functional pathways of immunocompetent cells. To extend our previous findings of the role of Period 2 in regulating splenocyte rhythms, we report that mice carrying a mutation in the Period 1 gene (Per1(-/-) mice), involved in the negative limb of the molecular clock, display significantly altered rhythms of cytokine (eg, interferon-γ) and cytolytic factors (eg, perforin and granzyme B) in splenic natural killer (NK) cells. Altered rhythms of NK cell immune factors were accompanied by changes in circadian expression of circadian clock genes, Bmal1 and Per2. In addition, Per1(-/-) circadian running-wheel activity rhythms remained rhythmic during constant darkness, although with a shortened free-running circadian period, suggesting primary involvement of peripheral molecular clocks. These findings indicate that the Per1 gene through NK cellular clocks modulates immune pathways.
Figures

Similar articles
-
Circadian oscillations of clock genes, cytolytic factors, and cytokines in rat NK cells.J Immunol. 2005 Jun 15;174(12):7618-24. doi: 10.4049/jimmunol.174.12.7618. J Immunol. 2005. PMID: 15944262
-
Evidence supporting a circadian control of natural killer cell function.Brain Behav Immun. 2006 Sep;20(5):469-76. doi: 10.1016/j.bbi.2005.10.002. Epub 2005 Nov 23. Brain Behav Immun. 2006. PMID: 16309885
-
Circadian rhythms of granzyme B, perforin, IFN-gamma, and NK cell cytolytic activity in the spleen: effects of chronic ethanol.J Immunol. 2004 Mar 1;172(5):2811-7. doi: 10.4049/jimmunol.172.5.2811. J Immunol. 2004. PMID: 14978081
-
[Molecular mechanisms of circadian clock functioning].Ukr Biokhim Zh (1999). 2011 May-Jun;83(3):5-24. Ukr Biokhim Zh (1999). 2011. PMID: 21888051 Review. Ukrainian.
-
Circadian modification network of a core clock driver BMAL1 to harmonize physiology from brain to peripheral tissues.Neurochem Int. 2018 Oct;119:11-16. doi: 10.1016/j.neuint.2017.12.013. Epub 2018 Jan 3. Neurochem Int. 2018. PMID: 29305918 Review.
Cited by
-
Circadian Rhythms in Immunity.Curr Allergy Asthma Rep. 2020 Jan 10;20(1):2. doi: 10.1007/s11882-020-0896-9. Curr Allergy Asthma Rep. 2020. PMID: 31925560 Free PMC article. Review.
-
Impact of noise exposure on the circadian clock in the auditory system.J Acoust Soc Am. 2019 Nov;146(5):3960. doi: 10.1121/1.5132290. J Acoust Soc Am. 2019. PMID: 31795664 Free PMC article. Review.
-
Circadian rhythms in leukocyte trafficking.Semin Immunopathol. 2014 Mar;36(2):149-62. doi: 10.1007/s00281-013-0414-4. Epub 2014 Jan 17. Semin Immunopathol. 2014. PMID: 24435096 Review.
-
Advances in the research of melatonin in autism spectrum disorders: literature review and new perspectives.Int J Mol Sci. 2013 Oct 14;14(10):20508-42. doi: 10.3390/ijms141020508. Int J Mol Sci. 2013. PMID: 24129182 Free PMC article. Review.
-
Circadian rhythms in solid organ transplantation.J Heart Lung Transplant. 2024 May;43(5):849-857. doi: 10.1016/j.healun.2024.01.017. Epub 2024 Feb 2. J Heart Lung Transplant. 2024. PMID: 38310995 Free PMC article. Review.
References
-
- Albrecht U. Timing to perfection: the biology of central and peripheral circadian clocks. Neuron. 2012;74(2):246–260. - PubMed
-
- Arjona A. Boyadjieva N. Sarkar DK. Circadian rhythms of granzyme B, perforin, IFNgamma, and NK cell cytolytic activity in the spleen: effects of chronic ethanol. J Immunol. 2004;172(5):2811–2817. - PubMed
-
- Arjona A. Sarkar DK. Circadian oscillations of clock genes, cytolytic factors, and cytokines in rat NK cells. J Immunol. 2005;174(12):7618–7624. - PubMed
-
- Arjona A. Sarkar DK. The circadian gene mPer2 regulates the daily rhythm of IFNgamma. J Interferon Cytokine Res. 2006a;26(9):645–649. - PubMed
-
- Arjona A. Sarkar DK. Evidence supporting a circadian control of natural killer cell function. Brain Behav Immun. 2006b;20(5):469–476. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

