Signal recognition initiates reorganization of the presequence translocase during protein import
- PMID: 23403928
- PMCID: PMC3604718
- DOI: 10.1038/emboj.2013.23
Signal recognition initiates reorganization of the presequence translocase during protein import
Abstract
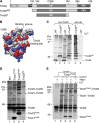
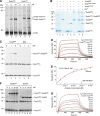
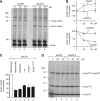
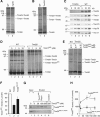
The mitochondrial presequence translocase interacts with presequence-containing precursors at the intermembrane space (IMS) side of the inner membrane to mediate their translocation into the matrix. Little is known as too how these matrix-targeting signals activate the translocase in order to initiate precursor transport. Therefore, we analysed how signal recognition by the presequence translocase initiates reorganization among Tim-proteins during import. Our analyses revealed that the presequence receptor Tim50 interacts with Tim21 in a signal-sensitive manner in a process that involves the IMS-domain of the Tim23 channel. The signal-driven release of Tim21 from Tim50 promotes recruitment of Pam17 and thus triggers formation of the motor-associated form of the TIM23 complex required for matrix transport.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures

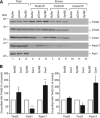

References
-
- Abe Y, Shodai T, Muto T, Mihara K, Torii H, Nishikawa S, Endo T, Kohda D (2000) Structural basis of presequence recognition by the mitochondrial protein import receptor Tom20. Cell 100: 551–560 - PubMed
-
- Bauer MF, Sirrenberg C, Neupert W, Brunner M (1996) Role of Tim23 as voltage sensor and presequence receptor in protein import into mitochondria. Cell 87: 33–41 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

