The RNA exosome complex central channel controls both exonuclease and endonuclease Dis3 activities in vivo and in vitro
- PMID: 23404585
- PMCID: PMC3616716
- DOI: 10.1093/nar/gkt060
The RNA exosome complex central channel controls both exonuclease and endonuclease Dis3 activities in vivo and in vitro
Abstract
The RNA exosome is an essential ribonuclease complex involved in RNA processing and decay. It consists of a 9-subunit catalytically inert ring composed of six RNase PH-like proteins forming a central channel and three cap subunits with KH/S1 domains located at the top. The yeast exosome catalytic activity is supplied by the Dis3 (also known as Rrp44) protein, which has both endo- and exoribonucleolytic activities and the nucleus-specific exonuclease Rrp6. In vitro studies suggest that substrates reach the Dis3 exonucleolytic active site following passage through the ring channel, but in vivo support is lacking. Here, we constructed an Rrp41 ring subunit mutant with a partially blocked channel that led to thermosensitivity and synthetic lethality with Rrp6 deletion. Rrp41 mutation caused accumulation of nuclear and cytoplasmic exosome substrates including the non-stop decay reporter, for which degradation is dependent on either endonucleolytic or exonucleolytic Dis3 activities. This suggests that the central channel also controls endonucleolytic activity. In vitro experiments performed using Chaetomium thermophilum exosomes reconstituted from recombinant subunits confirmed this notion. Finally, we analysed the impact of a lethal mutation of conserved basic residues in Rrp4 cap subunit and found that it inhibits digestion of single-stranded and structured RNA substrates.
Figures

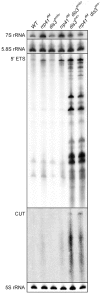






References
-
- Mitchell P, Petfalski E, Shevchenko A, Mann M, Tollervey D. The exosome: a conserved eukaryotic RNA processing complex containing multiple 3′→5′ exoribonucleases. Cell. 1997;91:457–466. - PubMed
-
- Tomecki R, Drazkowska K, Dziembowski A. Mechanisms of RNA degradation by the eukaryotic exosome. Chembiochem. 2010;11:938–945. - PubMed
-
- Isken O, Maquat LE. Quality control of eukaryotic mRNA: safeguarding cells from abnormal mRNA function. Genes Dev. 2007;21:1833–1856. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

