The PP6 phosphatase regulates ABI5 phosphorylation and abscisic acid signaling in Arabidopsis
- PMID: 23404889
- PMCID: PMC3608775
- DOI: 10.1105/tpc.112.105767
The PP6 phosphatase regulates ABI5 phosphorylation and abscisic acid signaling in Arabidopsis
Abstract
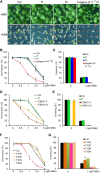
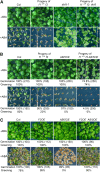
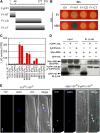
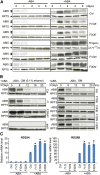
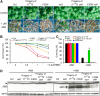
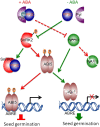
The basic Leucine zipper transcription factor ABSCISIC ACID INSENSITIVE5 (ABI5) is a key regulator of abscisic acid (ABA)-mediated seed germination and postgermination seedling growth. While a family of SUCROSE NONFERMENTING1-related protein kinase2s (SnRK2s) is responsible for ABA-induced phosphorylation and stabilization of ABI5, the phosphatase(s) responsible for dephosphorylating ABI5 is still unknown. Here, we demonstrate that mutations in FyPP1 (for Phytochrome-associated serine/threonine protein phosphatase1) and FyPP3, two homologous genes encoding the catalytic subunits of Ser/Thr PROTEIN PHOSPHATASE6 (PP6), cause an ABA hypersensitive phenotype in Arabidopsis thaliana, including ABA-mediated inhibition of seed germination and seedling growth. Conversely, overexpression of FyPP causes reduced sensitivity to ABA. The ABA hypersensitive phenotype of FyPP loss-of-function mutants is ABI5 dependent, and the amount of phosphorylated and total ABI5 proteins inversely correlates with the levels of FyPP proteins. Moreover, FyPP proteins physically interact with ABI5 in vitro and in vivo, and the strength of the interaction depends on the ABI5 phosphorylation status. In vitro phosphorylation assays show that FyPP proteins directly dephosphorylate ABI5. Furthermore, genetic and biochemical assays show that FyPP proteins act antagonistically with SnRK2 kinases to regulate ABI5 phosphorylation and ABA responses. Thus, Arabidopsis PP6 phosphatase regulates ABA signaling through dephosphorylation and destabilization of ABI5.
Figures


References
-
- Bracha-Drori K., Shichrur K., Katz A., Oliva M., Angelovici R., Yalovsky S., Ohad N. (2004). Detection of protein-protein interactions in plants using bimolecular fluorescence complementation. Plant J. 40: 419–427 - PubMed
-
- Busk P.K., Pagès M. (1998). Regulation of abscisic acid-induced transcription. Plant Mol. Biol. 37: 425–435 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

