Identification of cardiolipin binding sites on cytochrome c oxidase at the entrance of proton channels
- PMID: 23405277
- PMCID: PMC3570132
- DOI: 10.1038/srep01263
Identification of cardiolipin binding sites on cytochrome c oxidase at the entrance of proton channels
Erratum in
- Sci Rep. 2013;3:1343
Abstract
The respiratory chain or oxidative phosphorylation system (OxPhos) generates most of the chemical energy (ATP) used by our cells. The cytochrome c oxidase (CcO) is one of three protein complexes of OxPhos building up a proton gradient across the inner mitochondrial membrane, which is ultimately used by the ATP synthase to produce ATP. We present molecular dynamic simulations of CcO in a mimic of the mitochondrial membrane, and identify precise binding sites of cardiolipin (CL, signature phospholipid of mitochondria) on the protein surface. Two of these CL binding sites reveal pathways linking CLs to the entrance of the D and H proton channels across CcO. CLs being able to carry protons our results strongly support an involvement of CLs in the proton delivery machinery to CcO. The ubiquitous nature of CL interactions with the components of the OxPhos suggests that this delivery mechanism might extend to the other respiratory complexes.
Figures


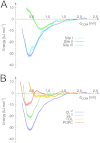



References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

