Structure and function of a cyanophage-encoded peptide deformylase
- PMID: 23407310
- PMCID: PMC3660681
- DOI: 10.1038/ismej.2013.4
Structure and function of a cyanophage-encoded peptide deformylase
Abstract
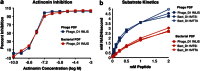
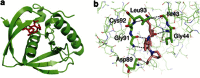
Bacteriophages encode auxiliary metabolic genes that support more efficient phage replication. For example, cyanophages carry several genes to maintain host photosynthesis throughout infection, shuttling the energy and reducing power generated away from carbon fixation and into anabolic pathways. Photodamage to the D1/D2 proteins at the core of photosystem II necessitates their continual replacement. Synthesis of functional proteins in bacteria requires co-translational removal of the N-terminal formyl group by a peptide deformylase (PDF). Analysis of marine metagenomes to identify phage-encoded homologs of known metabolic genes found that marine phages carry PDF genes, suggesting that their expression during infection might benefit phage replication. We identified a PDF homolog in the genome of Synechococcus cyanophage S-SSM7. Sequence analysis confirmed that it possesses the three absolutely conserved motifs that form the active site in PDF metalloproteases. Phylogenetic analysis placed it within the Type 1B subclass, most closely related to the Arabidopsis chloroplast PDF, but lacking the C-terminal α-helix characteristic of that group. PDF proteins from this phage and from Synechococcus elongatus were expressed and characterized. The phage PDF is the more active enzyme and deformylates the N-terminal tetrapeptides from D1 proteins more efficiently than those from ribosomal proteins. Solution of the X-ray/crystal structures of those two PDFs to 1.95 Å resolution revealed active sites identical to that of the Type 1B Arabidopsis chloroplast PDF. Taken together, these findings show that many cyanophages encode a PDF with a D1 substrate preference that adds to the repertoire of genes used by phages to maintain photosynthetic activities.
Figures


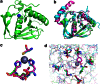
Similar articles
-
The C-terminal residue of phage Vp16 PDF, the smallest peptide deformylase, acts as an offset element locking the active conformation.Sci Rep. 2017 Sep 8;7(1):11041. doi: 10.1038/s41598-017-11329-3. Sci Rep. 2017. PMID: 28887476 Free PMC article.
-
Prevalence and evolution of core photosystem II genes in marine cyanobacterial viruses and their hosts.PLoS Biol. 2006 Jul;4(8):e234. doi: 10.1371/journal.pbio.0040234. PLoS Biol. 2006. PMID: 16802857 Free PMC article.
-
Peptide deformylases from Vibrio parahaemolyticus phage and bacteria display similar deformylase activity and inhibitor binding clefts.Biochim Biophys Acta Proteins Proteom. 2018 Feb;1866(2):348-355. doi: 10.1016/j.bbapap.2017.10.007. Epub 2017 Oct 31. Biochim Biophys Acta Proteins Proteom. 2018. PMID: 29101077
-
Cyanophage infection and photoinhibition in marine cyanobacteria.Res Microbiol. 2004 Nov;155(9):720-5. doi: 10.1016/j.resmic.2004.06.002. Res Microbiol. 2004. PMID: 15501648 Review.
-
Shedding new light on viral photosynthesis.Photosynth Res. 2015 Oct;126(1):71-97. doi: 10.1007/s11120-014-0057-x. Epub 2014 Nov 9. Photosynth Res. 2015. PMID: 25381655 Review.
Cited by
-
Marine and giant viruses as indicators of a marine microbial community in a riverine system.Microbiologyopen. 2016 Dec;5(6):1071-1084. doi: 10.1002/mbo3.392. Epub 2016 Aug 9. Microbiologyopen. 2016. PMID: 27506856 Free PMC article.
-
High coverage metabolomics analysis reveals phage-specific alterations to Pseudomonas aeruginosa physiology during infection.ISME J. 2016 Aug;10(8):1823-35. doi: 10.1038/ismej.2016.3. Epub 2016 Feb 16. ISME J. 2016. PMID: 26882266 Free PMC article.
-
Nitrogen sourcing during viral infection of marine cyanobacteria.Proc Natl Acad Sci U S A. 2019 Jul 30;116(31):15590-15595. doi: 10.1073/pnas.1901856116. Epub 2019 Jul 15. Proc Natl Acad Sci U S A. 2019. PMID: 31308237 Free PMC article.
-
Depth-stratified functional and taxonomic niche specialization in the 'core' and 'flexible' Pacific Ocean Virome.ISME J. 2015 Feb;9(2):472-84. doi: 10.1038/ismej.2014.143. Epub 2014 Aug 5. ISME J. 2015. PMID: 25093636 Free PMC article.
-
The C-terminal residue of phage Vp16 PDF, the smallest peptide deformylase, acts as an offset element locking the active conformation.Sci Rep. 2017 Sep 8;7(1):11041. doi: 10.1038/s41598-017-11329-3. Sci Rep. 2017. PMID: 28887476 Free PMC article.
References
-
- Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215:403–410. - PubMed
-
- Bingel-Erlenmeyer R, Kohler R, Kramer G, Arzu Sandikci S, Maier T, Schaffitzel C, et al. A peptide deformylase–ribosome complex reveals mechanism of nascent chain processing. Nature. 2008;452:108–111. - PubMed
-
- Bouzaidi-Tiali N, Giglione C, Bulliard Y, Pusnik M, Meinnel T, Schneider A. Type 3 peptide deformylases are required for oxidative phosphorylation in Trypanosoma brucei. Mol Microbiol. 2007;65:1218–1228. - PubMed
-
- Breitbart M, Thompson LR, Suttle CA, Sullivan MB. Exploring the vast diversity of marine viruses. Oceanography. 2007;20:135–139.
-
- Chen DZ, Patel DV, Hackbarth CJ, Wang W, Dreyer G, Young DC, et al. Actinonin, a naturally occurring antibacterial agent, is a potent deformylase inhibitor. Biochemistry. 2000;39:1256–1262. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

