Autophagy induction is a Tor- and Tp53-independent cell survival response in a zebrafish model of disrupted ribosome biogenesis
- PMID: 23408911
- PMCID: PMC3567153
- DOI: 10.1371/journal.pgen.1003279
Autophagy induction is a Tor- and Tp53-independent cell survival response in a zebrafish model of disrupted ribosome biogenesis
Abstract
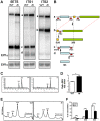
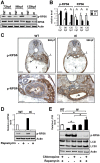
Ribosome biogenesis underpins cell growth and division. Disruptions in ribosome biogenesis and translation initiation are deleterious to development and underlie a spectrum of diseases known collectively as ribosomopathies. Here, we describe a novel zebrafish mutant, titania (tti(s450)), which harbours a recessive lethal mutation in pwp2h, a gene encoding a protein component of the small subunit processome. The biochemical impacts of this lesion are decreased production of mature 18S rRNA molecules, activation of Tp53, and impaired ribosome biogenesis. In tti(s450), the growth of the endodermal organs, eyes, brain, and craniofacial structures is severely arrested and autophagy is up-regulated, allowing intestinal epithelial cells to evade cell death. Inhibiting autophagy in tti(s450) larvae markedly reduces their lifespan. Somewhat surprisingly, autophagy induction in tti(s450) larvae is independent of the state of the Tor pathway and proceeds unabated in Tp53-mutant larvae. These data demonstrate that autophagy is a survival mechanism invoked in response to ribosomal stress. This response may be of relevance to therapeutic strategies aimed at killing cancer cells by targeting ribosome biogenesis. In certain contexts, these treatments may promote autophagy and contribute to cancer cells evading cell death.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures






References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

