Whole exome sequencing suggests much of non-BRCA1/BRCA2 familial breast cancer is due to moderate and low penetrance susceptibility alleles
- PMID: 23409019
- PMCID: PMC3568132
- DOI: 10.1371/journal.pone.0055681
Whole exome sequencing suggests much of non-BRCA1/BRCA2 familial breast cancer is due to moderate and low penetrance susceptibility alleles
Abstract
The identification of the two most prevalent susceptibility genes in breast cancer, BRCA1 and BRCA2, was the beginning of a sustained effort to uncover new genes explaining the missing heritability in this disease. Today, additional high, moderate and low penetrance genes have been identified in breast cancer, such as P53, PTEN, STK11, PALB2 or ATM, globally accounting for around 35 percent of the familial cases. In the present study we used massively parallel sequencing to analyze 7 BRCA1/BRCA2 negative families, each having at least 6 affected women with breast cancer (between 6 and 10) diagnosed under the age of 60 across generations. After extensive filtering, Sanger sequencing validation and co-segregation studies, variants were prioritized through either control-population studies, including up to 750 healthy individuals, or case-control assays comprising approximately 5300 samples. As a result, a known moderate susceptibility indel variant (CHEK2 1100delC) and a catalogue of 11 rare variants presenting signs of association with breast cancer were identified. All the affected genes are involved in important cellular mechanisms like DNA repair, cell proliferation and survival or cell cycle regulation. This study highlights the need to investigate the role of rare variants in familial cancer development by means of novel high throughput analysis strategies optimized for genetically heterogeneous scenarios. Even considering the intrinsic limitations of exome resequencing studies, our findings support the hypothesis that the majority of non-BRCA1/BRCA2 breast cancer families might be explained by the action of moderate and/or low penetrance susceptibility alleles.
Conflict of interest statement
Figures
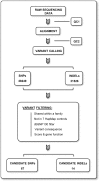
References
-
- Miki Y, Swensen J, Shattuck-Eidens D, Futreal PA, Harshman K, et al. (1994) A strong candidate for the breast and ovarian cancer susceptibility gene BRCA1. Science 266: 66–71. - PubMed
-
- Wooster R, Bignell G, Lancaster J, Swift S, Seal S, et al. (1995) Identification of the breast cancer susceptibility gene BRCA2. Nature 378: 789–792. - PubMed
-
- Borresen AL, Andersen TI, Garber J, Barbier-Piraux N, Thorlacius S, et al. (1992) Screening for germ line TP53 mutations in breast cancer patients. Cancer Res 52: 3234–3236. - PubMed
-
- Giardiello FM, Brensinger JD, Tersmette AC, Goodman SN, Petersen GM, et al. (2000) Very high risk of cancer in familial Peutz-Jeghers syndrome. Gastroenterology 119: 1447–1453. - PubMed
Publication types
MeSH terms
Grants and funding
- U01 CA069417/CA/NCI NIH HHS/United States
- U01 CA069638/CA/NCI NIH HHS/United States
- U01 CA69417/CA/NCI NIH HHS/United States
- U01 CA069467/CA/NCI NIH HHS/United States
- U01 CA069398/CA/NCI NIH HHS/United States
- U01 CA69638/CA/NCI NIH HHS/United States
- CA-06-503/CA/NCI NIH HHS/United States
- U01 CA069446/CA/NCI NIH HHS/United States
- U01 CA69467/CA/NCI NIH HHS/United States
- U01 CA069631/CA/NCI NIH HHS/United States
- U01 CA69398/CA/NCI NIH HHS/United States
- UL 2009-4388/PHS HHS/United States
- U01 CA69446/CA/NCI NIH HHS/United States
- U01 CA69631/CA/NCI NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

