Beyond secondary structure: primary-sequence determinants license pri-miRNA hairpins for processing
- PMID: 23415231
- PMCID: PMC3707628
- DOI: 10.1016/j.cell.2013.01.031
Beyond secondary structure: primary-sequence determinants license pri-miRNA hairpins for processing
Abstract
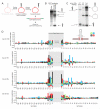
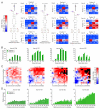
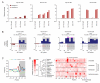
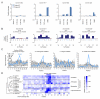
To use microRNAs to downregulate mRNA targets, cells must first process these ~22 nt RNAs from primary transcripts (pri-miRNAs). These transcripts form RNA hairpins important for processing, but additional determinants must distinguish pri-miRNAs from the many other hairpin-containing transcripts expressed in each cell. Illustrating the complexity of this recognition, we show that most Caenorhabditis elegans pri-miRNAs lack determinants required for processing in human cells. To find these determinants, we generated many variants of four human pri-miRNAs, sequenced millions that retained function, and compared them with the starting variants. Our results confirmed the importance of pairing in the stem and revealed three primary-sequence determinants, including an SRp20-binding motif (CNNC) found downstream of most pri-miRNA hairpins in bilaterian animals, but not in nematodes. Adding this and other determinants to C. elegans pri-miRNAs imparted efficient processing in human cells, thereby confirming the importance of primary-sequence determinants for distinguishing pri-miRNAs from other hairpin-containing transcripts.
Copyright © 2013 Elsevier Inc. All rights reserved.
Figures



References
-
- Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–297. - PubMed
-
- Bentwich I, Avniel A, Karov Y, Aharonov R, Gilad S, Barad O, Barzilai A, Einat P, Einav U, Meiri E, et al. Identification of hundreds of conserved and nonconserved human microRNAs. Nat Genet. 2005;37:766–770. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

