Nucleophosmin mutations in acute myeloid leukemia: a tale of protein unfolding and mislocalization
- PMID: 23436734
- PMCID: PMC3649256
- DOI: 10.1002/pro.2240
Nucleophosmin mutations in acute myeloid leukemia: a tale of protein unfolding and mislocalization
Abstract
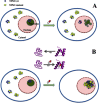
Nucleophosmin (NPM1) is an abundant, ubiquitously expressed protein mainly localized at nucleoli but continuously shuttling between nucleus and cytoplasm. NPM1 plays a role in several cellular functions, including ribosome biogenesis and export, centrosome duplication, chromatin remodeling, DNA repair, and response to stress stimuli. Much of the interest in this protein arises from its relevance in human malignancies. NPM1 is frequently overexpressed in solid tumors and is the target of several chromosomal translocations in hematologic neoplasms. Notably, NPM1 has been characterized as the most frequently mutated gene in acute myeloid leukemia (AML). Mutations alter the C-terminal DNA-binding domain of the protein and result in its aberrant nuclear export and stable cytosolic localization. In this review, we focus on the leukemia-associated NPM1 C-terminal domain and describe its structure, function, and the effect exerted by leukemic mutations. Finally, we discuss the possibility to target NPM1 for the treatment of cancer and, in particular, of AML patients with mutated NPM1 gene.
Copyright © 2013 The Protein Society.
Figures




References
-
- Kang YJ, Olson MO, Jones C, Busch H. Nucleolar phosphoproteins of normal rat liver and Novikoff hepatoma ascites cells. Cancer Res. 1975;35:1470–1475. - PubMed
-
- Borer RA, Lehner CF, Eppenberger HM, Nigg EA. Major nucleolar proteins shuttle between nucleus and cytoplasm. Cell. 1989;56:379–390. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

