Genome mining for methanobactins
- PMID: 23442874
- PMCID: PMC3621798
- DOI: 10.1186/1741-7007-11-17
Genome mining for methanobactins
Abstract
Background: Methanobactins (Mbns) are a family of copper-binding natural products involved in copper uptake by methanotrophic bacteria. The few Mbns that have been structurally characterized feature copper coordination by two nitrogen-containing heterocycles next to thioamide groups embedded in a peptidic backbone of varying composition. Mbns are proposed to derive from post-translational modification of ribosomally synthesized peptides, but only a few genes encoding potential precursor peptides have been identified. Moreover, the relevance of neighboring genes in these genomes has been unclear.
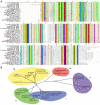
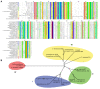
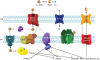
Results: The potential for Mbn production in a wider range of bacterial species was assessed by mining microbial genomes. Operons encoding Mbn-like precursor peptides, MbnAs, were identified in 16 new species, including both methanotrophs and, surprisingly, non-methanotrophs. Along with MbnA, the core of the operon is formed by two putative biosynthetic genes denoted MbnB and MbnC. The species can be divided into five groups on the basis of their MbnA and MbnB sequences and their operon compositions. Additional biosynthetic proteins, including aminotransferases, sulfotransferases and flavin adenine dinucleotide (FAD)-dependent oxidoreductases were also identified in some families. Beyond biosynthetic machinery, a conserved set of transporters was identified, including MATE multidrug exporters and TonB-dependent transporters. Additional proteins of interest include a di-heme cytochrome c peroxidase and a partner protein, the roles of which remain a mystery.
Conclusions: This study indicates that Mbn-like compounds may be more widespread than previously thought, but are not present in all methanotrophs. This distribution of species suggests a broader role in metal homeostasis. These data provide a link between precursor peptide sequence and Mbn structure, facilitating predictions of new Mbn structures and supporting a post-translational modification biosynthetic pathway. In addition, testable models for Mbn transport and for methanotrophic copper regulation have emerged. Given the unusual modifications observed in Mbns characterized thus far, understanding the roles of the putative biosynthetic proteins is likely to reveal novel pathways and chemistry.
Figures



References
-
- Semrau JD, Dispirito AA, Yoon S. Methanotrophs and copper. FEMS Microbiol Lett. 2010;34:496–531. - PubMed
-
- Jiang H, Chen Y, Jiang PX, Zhang C, Smith TJ, Murrell JC, Xing XH. Methanotrophs: Multifunctional bacteria with promising applications in environmental bioengineering. Biochem Eng J. 2010;49:277–288. doi: 10.1016/j.bej.2010.01.003. - DOI
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

