Defining the membrane-associated state of the PTEN tumor suppressor protein
- PMID: 23442912
- PMCID: PMC3566463
- DOI: 10.1016/j.bpj.2012.12.002
Defining the membrane-associated state of the PTEN tumor suppressor protein
Abstract
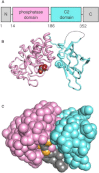
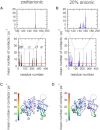
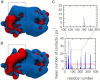
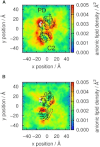
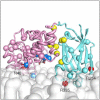
Phosphatase and tensin-homolog deleted on chromosome 10 (PTEN) is a tumor-suppressor protein that regulates phosphatidylinositol 3-kinase (PI3-K) signaling by binding to the plasma membrane and hydrolyzing the 3' phosphate from phosphatidylinositol (3,4,5)-trisphosphate (PI(3,4,5)P3) to form phosphatidylinositol (4,5)-bisphosphate (PI(4,5)P2). Several loss-of-function mutations in PTEN that impair lipid phosphatase activity and membrane binding are oncogenic, leading to the development of a variety of cancers, but information about the membrane-associated state of PTEN remains sparse. We have modeled a membrane-associated state of the truncated PTEN structure bound to PI(3,4,5)P3 via multiscale molecular dynamics simulations. We show that the location of the membrane-binding surface agrees with experimental observations and is robust to changes in lipid composition. The level of membrane interaction is substantially reduced in the phosphatase domain for the triple mutant R161E/K163E/K164E, in line with experimental results. We observe clustering of anionic lipids around the C2 domain in preference to the phosphatase domain, suggesting that the C2 domain is involved in nonspecific interactions with negatively charged lipid headgroups. Finally, our simulations suggest that the oncogenicity of the R335L mutation may be due to a reduction in the interaction of the mutant PTEN with anionic lipids.
Copyright © 2013 Biophysical Society. Published by Elsevier Inc. All rights reserved.
Figures


Similar articles
-
Interactions of the auxilin-1 PTEN-like domain with model membranes result in nanoclustering of phosphatidyl inositol phosphates.Biophys J. 2013 Jul 2;105(1):137-45. doi: 10.1016/j.bpj.2013.05.012. Biophys J. 2013. PMID: 23823232 Free PMC article.
-
Allosteric activation of PTEN phosphatase by phosphatidylinositol 4,5-bisphosphate.J Biol Chem. 2003 Sep 5;278(36):33617-20. doi: 10.1074/jbc.C300296200. Epub 2003 Jul 11. J Biol Chem. 2003. PMID: 12857747
-
Interactions of phosphatase and tensin homologue (PTEN) proteins with phosphatidylinositol phosphates: insights from molecular dynamics simulations of PTEN and voltage sensitive phosphatase.Biochemistry. 2014 Mar 25;53(11):1724-32. doi: 10.1021/bi5000299. Epub 2014 Mar 10. Biochemistry. 2014. PMID: 24588644 Free PMC article. Clinical Trial.
-
Membrane association of the PTEN tumor suppressor: neutron scattering and MD simulations reveal the structure of protein-membrane complexes.Methods. 2015 May;77-78:136-46. doi: 10.1016/j.ymeth.2014.10.014. Epub 2014 Oct 27. Methods. 2015. PMID: 25461777 Free PMC article. Review.
-
Biophysical methods for the characterization of PTEN/lipid bilayer interactions.Methods. 2015 May;77-78:125-35. doi: 10.1016/j.ymeth.2015.02.004. Epub 2015 Feb 16. Methods. 2015. PMID: 25697761 Free PMC article. Review.
Cited by
-
Binding of Ca2+-independent C2 domains to lipid membranes: A multi-scale molecular dynamics study.Structure. 2021 Oct 7;29(10):1200-1213.e2. doi: 10.1016/j.str.2021.05.011. Epub 2021 Jun 2. Structure. 2021. PMID: 34081910 Free PMC article.
-
A pathogenic role for germline PTEN variants which accumulate into the nucleus.Eur J Hum Genet. 2018 Aug;26(8):1180-1187. doi: 10.1038/s41431-018-0155-x. Epub 2018 Apr 30. Eur J Hum Genet. 2018. PMID: 29706633 Free PMC article.
-
Kinetics of PTEN-mediated PI(3,4,5)P3 hydrolysis on solid supported membranes.PLoS One. 2018 Feb 15;13(2):e0192667. doi: 10.1371/journal.pone.0192667. eCollection 2018. PLoS One. 2018. PMID: 29447222 Free PMC article.
-
Interactions of the auxilin-1 PTEN-like domain with model membranes result in nanoclustering of phosphatidyl inositol phosphates.Biophys J. 2013 Jul 2;105(1):137-45. doi: 10.1016/j.bpj.2013.05.012. Biophys J. 2013. PMID: 23823232 Free PMC article.
-
Aggregation of lipid-anchored full-length H-Ras in lipid bilayers: simulations with the MARTINI force field.PLoS One. 2013 Jul 26;8(7):e71018. doi: 10.1371/journal.pone.0071018. Print 2013. PLoS One. 2013. PMID: 23923044 Free PMC article.
References
-
- Lemmon M.A. Membrane recognition by phospholipid-binding domains. Nat. Rev. Mol. Cell Biol. 2008;9:99–111. - PubMed
-
- McLaughlin S. The electrostatic properties of membranes. Annu. Rev. Biophys. Biophys. Chem. 1989;18:113–136. - PubMed
-
- McLaughlin S., Wang J., Murray D. PIP2 and proteins: interactions, organization, and information flow. Annu. Rev. Biophys. Biomol. Struct. 2002;31:151–175. - PubMed
-
- McLaughlin S., Murray D. Plasma membrane phosphoinositide organization by protein electrostatics. Nature. 2005;438:605–611. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- WT_/Wellcome Trust/United Kingdom
- BBS/B/16011/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- BEP17032/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- B19456/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- MRC_/Medical Research Council/United Kingdom
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

