The influence of clan structure on the genetic variation in a single Ghanaian village
- PMID: 23443025
- PMCID: PMC3778349
- DOI: 10.1038/ejhg.2013.12
The influence of clan structure on the genetic variation in a single Ghanaian village
Abstract
Socioeconomic and cultural factors are thought to have an important role in influencing human population genetic structure. To explain such population structure differences, most studies analyse genetic differences among widely dispersed human populations. In contrast, we have studied the genetic structure of an ethnic group occupying a single village in north-eastern Ghana. We found a markedly skewed male population substructure because of an almost complete lack of male gene flow among Bimoba clans in this village. We also observed a deep male substructure within one of the clans in this village. Among all males, we observed only three Y-single-nucleotide polymorphism (SNP) haplogroups: E1b1a*-M2, E1b1a7a*-U174 and E1b1a8a*-U209, P277, P278. In contrast to the marked Y-chromosomal substructure, mitochondrial DNA HVS-1 sequence variation and autosomal short-tandem repeats variation patterns indicate high genetic diversities and a virtually random female-mediated gene flow among clans. On the extreme micro-geographical scale of this single Bimoba village, correspondence between the Y-chromosome lineages and clan membership could be due to the combined effects of the strict patrilocal and patrilineal structure. If translated to larger geographic scales, our results would imply that the extent of variation in uniparentally inherited genetic markers, which are typically associated with historical migration on a continental scale, could equally likely be the result of many small and different cumulative effects of social factors such as clan membership that act at a local scale. Such local scale effects should therefore be considered in genetic studies, especially those that use uniparental markers, before making inferences about human history at large.
Figures
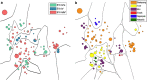
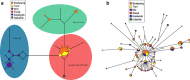
References
-
- Cavalli-Sforza LL, Menozzi P, Piazza A. The History and Geography of Human Genes. Princeton Yniversity Press; 1994.
-
- Curran SR, Agardy T. Common property systems, migration, and coastal ecosystems. J Human Environ. 2002;31:303–305. - PubMed
-
- Barbujani G, Colonna V. Human genome diversity: frequently asked questions. Trends Genet. 2010;26:285–295. - PubMed
-
- Wollstein A, Lao O, Becker C, et al. Demographic history of Oceania inferred from genome-wide data. Curr Biol. 2010;20:1983–1992. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

