Expression, purification and biochemical characterization of Schizosaccharomyces pombe Mcm4, 6 and 7
- PMID: 23444842
- PMCID: PMC3605359
- DOI: 10.1186/1471-2091-14-5
Expression, purification and biochemical characterization of Schizosaccharomyces pombe Mcm4, 6 and 7
Abstract
Background: The hetero-hexamer of the eukaryotic minichromosome maintenance (MCM) proteins plays an essential role in replication of genomic DNA. The ring-shaped Mcm2-7 hexamers comprising one of each subunit show helicase activity in vitro, and form double-hexamers on DNA. The Mcm4/6/7 also forms a hexameric complex with helicase activity in vitro.
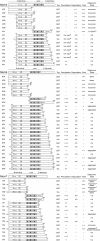
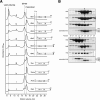
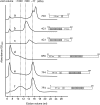
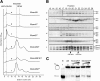
Results: We used an Escherichiai coli expression system to express various domains of Schizosaccharomyces pombe Mcm4, 6 and 7 in order to characterize their domain structure, oligomeric states, and possible inter-/intra-subunit interactions. We also successfully employed a co-expression system to express Mcm4/6/7 at the same time in Escherichiai coli, and have purified functional Mcm4/6/7 complex in a hexameric state in high yield and purity, providing a means for generating large quantity of proteins for future structural and biochemical studies.
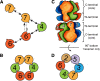
Conclusions: Based on our results and those of others, models were proposed for the subunit arrangement and architecture of both the Mcm4/6/7 hexamer and the Mcm2-7 double-hexamer.
Figures


Similar articles
-
Roles of Mcm7 and Mcm4 subunits in the DNA helicase activity of the mouse Mcm4/6/7 complex.J Biol Chem. 2002 Nov 8;277(45):42471-9. doi: 10.1074/jbc.M205769200. Epub 2002 Aug 30. J Biol Chem. 2002. PMID: 12207017
-
The effects of oligomerization on Saccharomyces cerevisiae Mcm4/6/7 function.BMC Biochem. 2010 Sep 22;11:37. doi: 10.1186/1471-2091-11-37. BMC Biochem. 2010. PMID: 20860810 Free PMC article.
-
Mcm4 C-terminal domain of MCM helicase prevents excessive formation of single-stranded DNA at stalled replication forks.Proc Natl Acad Sci U S A. 2008 Sep 2;105(35):12973-8. doi: 10.1073/pnas.0805307105. Epub 2008 Aug 27. Proc Natl Acad Sci U S A. 2008. PMID: 18753627 Free PMC article.
-
Reconstitution of the Mcm2-7p heterohexamer, subunit arrangement, and ATP site architecture.J Biol Chem. 2003 Feb 14;278(7):4491-9. doi: 10.1074/jbc.M210511200. Epub 2002 Dec 11. J Biol Chem. 2003. PMID: 12480933
-
Cdt1 forms a complex with the minichromosome maintenance protein (MCM) and activates its helicase activity.J Biol Chem. 2008 Sep 5;283(36):24469-77. doi: 10.1074/jbc.M803212200. Epub 2008 Jul 7. J Biol Chem. 2008. PMID: 18606811 Free PMC article.
Cited by
-
Arabidopsis thaliana MCM3 single subunit of MCM2-7 complex functions as 3' to 5' DNA helicase.Protoplasma. 2016 Mar;253(2):467-75. doi: 10.1007/s00709-015-0825-2. Epub 2015 May 6. Protoplasma. 2016. PMID: 25944245
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

