Transcription factor nuclear factor erythroid-2 mediates expression of the cytokine interleukin 8, a known predictor of inferior outcome in patients with myeloproliferative neoplasms
- PMID: 23445878
- PMCID: PMC3696611
- DOI: 10.3324/haematol.2012.071183
Transcription factor nuclear factor erythroid-2 mediates expression of the cytokine interleukin 8, a known predictor of inferior outcome in patients with myeloproliferative neoplasms
Abstract
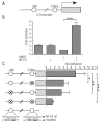
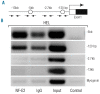
The transcription factor nuclear factor erythroid-2 is over-expressed in patients with myeloproliferative neoplasms irrespective of the presence of the JAK2(V617F) mutation. Our transgenic mouse model over-expressing nuclear factor erythroid-2, which recapitulates many features of myeloproliferative neoplasms including transformation to acute myeloid leukemia, clearly implicates this transcription factor in the pathophysiology of myeloproliferative neoplasms. Because the targets mediating nuclear factor erythroid-2 effects are not well characterized, we conducted microarray analysis of CD34(+) cells lentivirally transduced to over-express nuclear factor erythroid-2 or to silence this transcription factor via shRNA, in order to identify novel target genes. Here, we report that the cytokine interleukin 8 is a novel target gene. Nuclear factor erythroid-2 directly binds the interleukin 8 promoter in vivo, and these binding sites are required for promoter activity. Serum levels of interleukin 8 are known to be elevated in both polycythemia vera and primary myelofibrosis patients. Recently, increased interleukin 8 levels have been shown to be predictive of inferior survival in primary myelofibrosis patients in multivariate analysis. Therefore, one of the mechanisms by which nuclear factor erythroid-2 contributes to myeloproliferative neoplasm pathology may be increased interleukin 8 expression.
Figures



References
-
- James C, Ugo V, Le Couedic JP, Staerk J, Delhommeau F, Lacout C, et al. A unique clonal JAK2 mutation leading to constitutive signalling causes polycythaemia vera. Nature. 2005;434(7037):1144–8 - PubMed
-
- Cario H, Goerttler PS, Steimle C, Levine RL, Pahl HL. The JAK2V617F mutation is acquired secondary to the predisposing alteration in familial polycythaemia vera. Br J Haematol. 2005;130(5):800–1 - PubMed
-
- Bellanné-Chantelot C, Chaumarel I, Labopin M, Bellanger F, Barbu V, De Toma C, et al. Genetic and clinical implications of the Val617Phe JAK2 mutation in 72 families with myeloproliferative disorders. Blood. 2006;108(1):346–52 - PubMed
-
- Kralovics R, Teo SS, Li S, Theocharides A, Buser AS, Tichelli A, et al. Acquisition of the V617F mutation of JAK2 is a late genetic event in a subset of patients with myeloproliferative disorders. Blood. 2006;108(4): 1377–80 - PubMed
-
- Nussenzveig RH, Swierczek SI, Jelinek J, Gaikwad A, Liu E, Verstovsek S, et al. Polycythemia vera is not initiated by JAK2V617F mutation. Exp Hematol. 2007; 35(1):32–8 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

