Mapping and analysis of phosphorylation sites: a quick guide for cell biologists
- PMID: 23447708
- PMCID: PMC3583658
- DOI: 10.1091/mbc.E12-09-0677
Mapping and analysis of phosphorylation sites: a quick guide for cell biologists
Abstract
A mechanistic understanding of signaling networks requires identification and analysis of phosphorylation sites. Mass spectrometry offers a rapid and highly sensitive approach to mapping phosphorylation sites. However, mass spectrometry has significant limitations that must be considered when planning to carry out phosphorylation-site mapping. Here we provide an overview of key information that should be taken into consideration before beginning phosphorylation-site analysis, as well as a step-by-step guide for carrying out successful experiments.
Figures

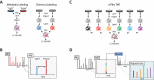
References
-
- Beausoleil SA, Villen J, Gerber SA, Rush J, Gygi SP. A probability-based approach for high-throughput protein phosphorylation analysis and site localization. Nat Biotechnol. 2006;24:1285–1292. - PubMed
-
- Boersema PJ, Raijmakers R, Lemeer S, Mohammed S, Heck AJ. Multiplex peptide stable isotope dimethyl labeling for quantitative proteomics. Nat Protoc. 2009;4:484–494. - PubMed
-
- Boyle WJ, van der Geer P, Hunter T. Phosphopeptide mapping and phosphoamino acid analysis by two-dimensional separation on thin-layer cellulose plates. Methods Enzymol. 1991;201:110–149. - PubMed
-
- Choi KY, Satterberg B, Lyons DM, Elion EA. Ste5 tethers multiple protein kinases in the MAP kinase cascade required for mating in S. cerevisiae. Cell. 1994;78:499–512. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

