pH-sensitive residues in the p19 RNA silencing suppressor protein from carnation Italian ringspot virus affect siRNA binding stability
- PMID: 23450521
- PMCID: PMC3649261
- DOI: 10.1002/pro.2243
pH-sensitive residues in the p19 RNA silencing suppressor protein from carnation Italian ringspot virus affect siRNA binding stability
Abstract
Tombusviruses, such as Carnation Italian ringspot virus (CIRV), encode a protein homodimer called p19 that is capable of suppressing RNA silencing in their infected hosts by binding to and sequestering short-interfering RNA (siRNA) away from the RNA silencing pathway. P19 binding stability has been shown to be sensitive to changes in pH but the specific amino acid residues involved have remained unclear. Using constant pH molecular dynamics simulations, we have identified key pH-dependent residues that affect CIRV p19-siRNA binding stability at various pH ranges based on calculated changes in the free energy contribution from each titratable residue. At high pH, the deprotonation of Lys60, Lys67, Lys71, and Cys134 has the largest effect on the binding stability. Similarly, deprotonation of several acidic residues (Asp9, Glu12, Asp20, Glu35, and/or Glu41) at low pH results in a decrease in binding stability. At neutral pH, residues Glu17 and His132 provide a small increase in the binding stability and we find that the optimal pH range for siRNA binding is between 7.0 and 10.0. Overall, our findings further inform recent experiments and are in excellent agreement with data on the pH-dependent binding profile.
Copyright © 2013 The Protein Society.
Figures
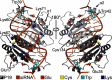
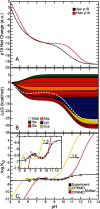
 values increase binding stability (i.e., favors holo form) while negative
values increase binding stability (i.e., favors holo form) while negative  values destabilizes binding (i.e., favors apo form). (C) pH-dependent dissociation constant profiles compared with experiment. CPHMDshifted (red) is identical to CPHMD (yellow) except that it is shifted to the right by 1.5 pH units for comparison with experiment (black).
values destabilizes binding (i.e., favors apo form). (C) pH-dependent dissociation constant profiles compared with experiment. CPHMDshifted (red) is identical to CPHMD (yellow) except that it is shifted to the right by 1.5 pH units for comparison with experiment (black).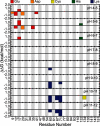
 and
and  , respectively, for a given titratable residue. Only residues with
, respectively, for a given titratable residue. Only residues with  are shown where
are shown where  is the Boltzmann constant and T is the temperature (298 K).
is the Boltzmann constant and T is the temperature (298 K).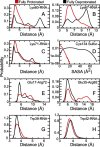
Similar articles
-
Effects of pH and salt concentration on the siRNA binding activity of the RNA silencing suppressor protein p19.FEBS Lett. 2007 Jun 26;581(16):3051-6. doi: 10.1016/j.febslet.2007.05.064. Epub 2007 Jun 4. FEBS Lett. 2007. PMID: 17559839
-
Cysteine residues of Carnation Italian Ringspot virus p19 suppressor of RNA silencing maintain global structural integrity and stability for siRNA binding.Biochim Biophys Acta. 2009 Aug;1794(8):1197-203. doi: 10.1016/j.bbapap.2009.03.012. Epub 2009 Mar 31. Biochim Biophys Acta. 2009. PMID: 19341824
-
Protein mediated miRNA detection and siRNA enrichment using p19.Biotechniques. 2010 Jun;48(6):xvii-xxiii. doi: 10.2144/000113364. Biotechniques. 2010. PMID: 20569217
-
Plant RNAi: How a viral silencing suppressor inactivates siRNA.Curr Biol. 2004 Mar 9;14(5):R198-200. doi: 10.1016/j.cub.2004.02.021. Curr Biol. 2004. PMID: 15028237 Review.
-
Studying the RNA silencing pathway with the p19 protein.FEBS Lett. 2013 Apr 17;587(8):1198-205. doi: 10.1016/j.febslet.2013.01.036. Epub 2013 Jan 29. FEBS Lett. 2013. PMID: 23376479 Review.
Cited by
-
Constant pH molecular dynamics of proteins in explicit solvent with proton tautomerism.Proteins. 2014 Jul;82(7):1319-31. doi: 10.1002/prot.24499. Epub 2014 Jan 15. Proteins. 2014. PMID: 24375620 Free PMC article.
-
Recent development and application of constant pH molecular dynamics.Mol Simul. 2014 Jan 1;40(10-11):830-838. doi: 10.1080/08927022.2014.907492. Mol Simul. 2014. PMID: 25309035 Free PMC article.
-
Uncovering pH-dependent transient states of proteins with buried ionizable residues.J Am Chem Soc. 2014 Jun 18;136(24):8496-9. doi: 10.1021/ja5012564. Epub 2014 Jun 3. J Am Chem Soc. 2014. PMID: 24842060 Free PMC article.
-
Membrane environment modulates the pKa values of transmembrane helices.J Phys Chem B. 2015 Apr 2;119(13):4601-7. doi: 10.1021/acs.jpcb.5b00289. Epub 2015 Mar 22. J Phys Chem B. 2015. PMID: 25734901 Free PMC article.
-
CHARMM-GUI 10 years for biomolecular modeling and simulation.J Comput Chem. 2017 Jun 5;38(15):1114-1124. doi: 10.1002/jcc.24660. Epub 2016 Nov 14. J Comput Chem. 2017. PMID: 27862047 Free PMC article. Review.
References
-
- Fire A, Xu SQ, Montgomery MK, Kostas SA, Driver SE, Mello CC. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature. 1998;391:806–811. - PubMed
-
- Hannon GJ. Rna interference. Nature. 2002;418:244–251. - PubMed
-
- Voinnet O. RNA silencing as a plant immune system against viruses. Trends Genet. 2001;17:449–459. - PubMed
-
- Bernstein E, Caudy AA, Hammond SM, Hannon GJ. Role for a bidentate ribonuclease in the initiation step of RNA interference. Nature. 2001;409:363–366. - PubMed
-
- Hutvagner G, Simard MJ. Argonaute proteins: key players in RNA silencing. Nat Rev Mol Cell Bio. 2008;9:22–32. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

