Multi-disciplinary methods to define RNA-protein interactions and regulatory networks
- PMID: 23453689
- PMCID: PMC3624891
- DOI: 10.1016/j.gde.2013.01.003
Multi-disciplinary methods to define RNA-protein interactions and regulatory networks
Abstract
The advent of high-throughput technologies including deep-sequencing and protein mass spectrometry is facilitating the acquisition of large and precise data sets toward the definition of post-transcriptional regulatory networks. While early studies that investigated specific RNA-protein interactions in isolation laid the foundation for our understanding of the existence of molecular machines to assemble and process RNAs, there is a more recent appreciation of the importance of individual RNA-protein interactions that contribute to post-transcriptional gene regulation. The multitude of RNA-binding proteins (RBPs) and their many RNA targets has only been captured experimentally in recent times. In this review, we will examine current multidisciplinary approaches toward elucidating RNA-protein networks and their regulation.
Copyright © 2013 Elsevier Ltd. All rights reserved.
Figures

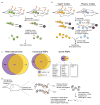
References
-
- Licatalosi DD, Mele A, Fak JJ, Ule J, Kayikci M, Chi SW, Clark TA, Schweitzer AC, Blume JE, Wang X, et al. HITS-CLIP yields genome-wide insights into brain alternative RNA processing. Nature. 2008;456:464–469. The first published report of UV RNA-protein crosslinking methods combined with next generation RNA sequencing methods to obtain genome-wide binding sites of RNA binding proteins. - PMC - PubMed
-
- Hafner M, Landthaler M, Burger L, Khorshid M, Hausser J, Berninger P, Rothballer A, Ascano PhDM, Jr, Jungkamp A-C, Munschauer M, et al. Transcriptome-wide Identification of RNA-Binding Protein and MicroRNA Target Sites by PAR-CLIP. Cell. 2010;141:129–141. Identification of genome-wide RNA targets by PAR-CLIP for the RNA binding proteins PUM2, QKI, IGF2BP, AGO(1–4), and TNRC6(A–C) - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

