The human leukocyte antigen-presented ligandome of B lymphocytes
- PMID: 23481700
- PMCID: PMC3708169
- DOI: 10.1074/mcp.M112.024810
The human leukocyte antigen-presented ligandome of B lymphocytes
Abstract
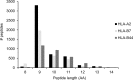
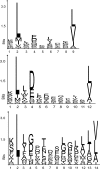
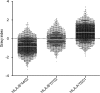
Peptides presented by human leukocyte antigen (HLA) molecules on the cell surface play a crucial role in adaptive immunology, mediating the communication between T cells and antigen presenting cells. Knowledge of these peptides is of pivotal importance in fundamental studies of T cell action and in cellular immunotherapy and transplantation. In this paper we present the in-depth identification and relative quantification of 14,500 peptide ligands constituting the HLA ligandome of B cells. This large number of identified ligands provides general insight into the presented peptide repertoire and antigen presentation. Our uniquely large set of HLA ligands allowed us to characterize in detail the peptides constituting the ligandome in terms of relative abundance, peptide length distribution, physicochemical properties, binding affinity to the HLA molecule, and presence of post-translational modifications. The presented B-lymphocyte ligandome is shown to be a rich source of information by the presence of minor histocompatibility antigens, virus-derived epitopes, and post-translationally modified HLA ligands, and it can be a good starting point for solving a wealth of specific immunological questions. These HLA ligands can form the basis for reversed immunology approaches to identify T cell epitopes based not on in silico predictions but on the bona fide eluted HLA ligandome.
Figures






References
-
- Kenter G. G., Welters M. J., Valentijn A. R., Lowik M. J., Berends-van der Meer D. M., Vloon A. P., Essahsah F., Fathers L. M., Offringa R., Drijfhout J. W., Wafelman A. R., Oostendorp J., Fleuren G. J., van der Burg S. H., Melief C. J. (2009) Vaccination against HPV-16 oncoproteins for vulvar intraepithelial neoplasia. N. Engl. J. Med. 361, 1838–1847 - PubMed
-
- Falkenburg J. H., Marijt W. A., Heemskerk M. H., Willemze R. (2002) Minor histocompatibility antigens as targets of graft-versus-leukemia reactions. Curr. Opin. Hematol. 9, 497–502 - PubMed
-
- Yewdell J. W., Reits E., Neefjes J. (2003) Making sense of mass destruction: quantitating MHC class I antigen presentation. Nat. Rev. Immunol. 3, 952–961 - PubMed
-
- Kessler J. H., Khan S., Seifert U., Le Gall S., Chow K. M., Paschen A., Bres-Vloemans S. A., de Ru A., van Montfoort N., Franken K. L., Benckhuijsen W. E., Brooks J. M., van Hall T., Ray K., Mulder A., Doxiadis I. I., van Swieten P. F., Overkleeft H. S., Prat A., Tomkinson B., Neefjes J., Kloetzel P. M., Rodgers D. W., Hersh L. B., Drijfhout J. W., van Veelen P. A., Ossendorp F., Melief C. J. (2011) Antigen processing by nardilysin and thimet oligopeptidase generates cytotoxic T cell epitopes. Nat. Immunol. 12, 45–53 - PubMed
-
- Vigneron N., Stroobant V., Chapiro J., Ooms A., Degiovanni G., Morel S., van der Bruggen P., Boon T., Van den Eynde B. J. (2004) An antigenic peptide produced by peptide splicing in the proteasome. Science 304, 587–590 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

