Sensitive cell-based assay for determination of human immunodeficiency virus type 1 coreceptor tropism
- PMID: 23486708
- PMCID: PMC3647936
- DOI: 10.1128/JCM.00092-13
Sensitive cell-based assay for determination of human immunodeficiency virus type 1 coreceptor tropism
Abstract
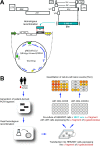
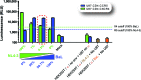
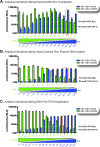
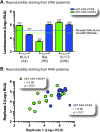
CCR5 antagonists are a powerful new class of antiretroviral drugs that require a companion assay to evaluate the presence of CXCR4-tropic (non-R5) viruses prior to use in human immunodeficiency virus (HIV)-infected individuals. In this study, we have developed, characterized, verified, and prevalidated a novel phenotypic test to determine HIV-1 coreceptor tropism (VERITROP) based on a sensitive cell-to-cell fusion assay. A proprietary vector was constructed containing a near-full-length HIV-1 genome with the yeast uracil biosynthesis (URA3) gene replacing the HIV-1 env coding sequence. Patient-derived HIV-1 PCR products were introduced by homologous recombination using an innovative yeast-based cloning strategy. The env-expressing vectors were then used in a cell-to-cell fusion assay to determine the presence of R5 and/or non-R5 HIV-1 variants within the viral population. Results were compared with (i) the original version of Trofile (Monogram Biosciences, San Francisco, CA), (ii) population sequencing, and (iii) 454 pyrosequencing, with the genotypic data analyzed using several bioinformatics tools, i.e., the 11/24/25 rule, Geno2Pheno (2% to 5.75%, 3.5%, or 10% false-positive rate [FPR]), and webPSSM. VERITROP consistently detected minority non-R5 variants from clinical specimens, with an analytical sensitivity of 0.3%, with viral loads of ≥1,000 copies/ml, and from B and non-B subtypes. In a pilot study, a 73.7% (56/76) concordance was observed with the original Trofile assay, with 19 of the 20 discordant results corresponding to non-R5 variants detected using VERITROP and not by the original Trofile assay. The degree of concordance of VERITROP and Trofile with population and deep sequencing results depended on the algorithm used to determine HIV-1 coreceptor tropism. Overall, VERITROP showed better concordance with deep sequencing/Geno2Pheno at a 0.3% detection threshold (67%), whereas Trofile matched better with population sequencing (79%). However, 454 sequencing using Geno2Pheno at a 10% FPR and 0.3% threshold and VERITROP more accurately predicted the success of a maraviroc-based regimen. In conclusion, VERITROP may promote the development of new HIV coreceptor antagonists and aid in the treatment and management of HIV-infected individuals prior to and/or during treatment with this class of drugs.
Figures
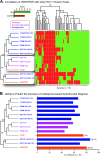
References
-
- Dalgleish AG, Beverley PC, Clapham PR, Crawford DH, Greaves MF, Weiss RA. 1984. The CD4 (T4) antigen is an essential component of the receptor for the AIDS retrovirus. Nature 312:763–767 - PubMed
-
- Klatzmann D, Champagne E, Chamaret S, Gruest J, Guetard D, Hercend T, Gluckman JC, Montagnier L. 1984. T-lymphocyte T4 molecule behaves as the receptor for human retrovirus LAV. Nature 312:767–768 - PubMed
-
- Dragic T, Litwin V, Allaway GP, Martin SR, Huang Y, Nagashima KA, Cayanan C, Maddon PJ, Koup RA, Moore JP, Paxton WA. 1996. HIV-1 entry into CD4+ cells is mediated by the chemokine receptor CC- CKR-5. Nature 381:667–673 - PubMed
-
- Feng Y, Broder CC, Kennedy PE, Berger EA. 1996. HIV-1 entry cofactor: functional cDNA cloning of a seven-transmembrane, G protein-coupled receptor. Science 272:872–877 - PubMed
-
- Deng H, Liu R, Ellmeier W, Choe S, Unutmaz D, Burkhart M, Di Marzio P, Marmon S, Sutton RE, Hill CM, Davis CB, Peiper SC, Schall TJ, Littman DR, Landau NR. 1996. Identification of a major co-receptor for primary isolates of HIV-1. Nature 381:661–666 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

