Determinants of human cyclin B1 association with mitotic chromosomes
- PMID: 23505570
- PMCID: PMC3594322
- DOI: 10.1371/journal.pone.0059169
Determinants of human cyclin B1 association with mitotic chromosomes
Abstract
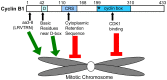
Cyclin B1-CDK1 activity is essential for mitotic entry, but questions remain regarding how the activity of this kinase is spatially regulated. Previous studies showed that the cyclin B1 subunit localizes to several compartments of a mitotic cell, including the centrosomes, mitotic spindle, kinetochores and chromosomes via distinct sequence elements. Mitotic chromosome association occurs through the unstructured N-terminal domain of cyclin B1 and is independent of CDK1 binding. Here, we use live cell imaging of human cyclin B1 fused to GFP to precisely define the sequence elements within cyclin B1 that mediate its association with condensed mitotic chromosomes. We find that a short, evolutionarily conserved N-terminal motif is required for cyclin B1 to localize to mitotic chromosomes. We further reveal a role for arginine residues within and near the destruction box sequence in the chromosome association of cyclin B1. Additionally, our data suggest that sequences further downstream in cyclin B1, such as the cytoplasmic retention sequence and the cyclin box, may negatively modulate chromosome association. Because multiple basic residues are required for cyclin B1 association with mitotic chromosomes, electrostatic interactions with DNA may facilitate cyclin B1 localization to chromosomes.
Conflict of interest statement
Figures

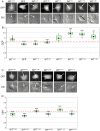
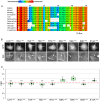
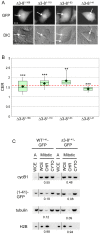
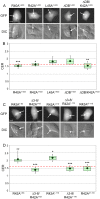
Similar articles
-
Phosphorylation of Mcl-1 by CDK1-cyclin B1 initiates its Cdc20-dependent destruction during mitotic arrest.EMBO J. 2010 Jul 21;29(14):2407-20. doi: 10.1038/emboj.2010.112. Epub 2010 Jun 4. EMBO J. 2010. PMID: 20526282 Free PMC article.
-
The APC/C recruits cyclin B1-Cdk1-Cks in prometaphase before D box recognition to control mitotic exit.J Cell Biol. 2010 Aug 23;190(4):587-602. doi: 10.1083/jcb.200912084. J Cell Biol. 2010. PMID: 20733055 Free PMC article.
-
Distinct sequence elements of cyclin B1 promote localization to chromatin, centrosomes, and kinetochores during mitosis.Mol Biol Cell. 2007 Dec;18(12):4847-58. doi: 10.1091/mbc.e06-06-0539. Epub 2007 Sep 19. Mol Biol Cell. 2007. PMID: 17881737 Free PMC article.
-
Two distinct controls of mitotic cdk1/cyclin B1 activity requisite for cell growth prior to cell division.Cell Cycle. 2007 Jun 15;6(12):1419-25. Epub 2007 May 7. Cell Cycle. 2007. PMID: 17592253 Review.
-
'... The end of the beginning': cdk1 thresholds and exit from mitosis.Cell Cycle. 2007 Jun 15;6(12):1408-11. Epub 2007 Apr 27. Cell Cycle. 2007. PMID: 17581279 Review.
Cited by
-
Spatial control of the APC/C ensures the rapid degradation of cyclin B1.EMBO J. 2024 Oct;43(19):4324-4355. doi: 10.1038/s44318-024-00194-2. Epub 2024 Aug 14. EMBO J. 2024. PMID: 39143240 Free PMC article.
-
Cyclin B1 destruction box-mediated protein instability: the enhanced sensitivity of fluorescent-protein-based reporter gene system.Biomed Res Int. 2013;2013:732307. doi: 10.1155/2013/732307. Epub 2013 Dec 15. Biomed Res Int. 2013. PMID: 24416725 Free PMC article.
-
The spindle and kinetochore-associated (Ska) complex enhances binding of the anaphase-promoting complex/cyclosome (APC/C) to chromosomes and promotes mitotic exit.Mol Biol Cell. 2014 Mar;25(5):594-605. doi: 10.1091/mbc.E13-07-0421. Epub 2014 Jan 8. Mol Biol Cell. 2014. PMID: 24403607 Free PMC article.
-
Initial characterization of gap phase introduction in every cell cycle of C. elegans embryogenesis.Front Cell Dev Biol. 2022 Oct 25;10:978962. doi: 10.3389/fcell.2022.978962. eCollection 2022. Front Cell Dev Biol. 2022. PMID: 36393848 Free PMC article.
-
The role of CDC25C in cell cycle regulation and clinical cancer therapy: a systematic review.Cancer Cell Int. 2020 Jun 3;20:213. doi: 10.1186/s12935-020-01304-w. eCollection 2020. Cancer Cell Int. 2020. PMID: 32518522 Free PMC article. Review.
References
-
- Morgan DO (2007) The Cell Cycle: Principles of Control; Lawrence E, editor. Corby: Oxford University Press. 297 p.
-
- Nigg EA (2001) Mitotic kinases as regulators of cell division and its checkpoints. Nat Rev Mol Cell Biol 2: 21–32. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

