Protein dynamics and the enzymatic reaction coordinate
- PMID: 23508766
- PMCID: PMC4346281
- DOI: 10.1007/128_2012_412
Protein dynamics and the enzymatic reaction coordinate
Abstract
This chapter discusses progress over the past 15 years in understanding the role of protein dynamics in enzymatically catalyzed chemical reactions. Research has shown that protein motion on all timescales from femtoseconds to milliseconds can contribute to function, and in particular in some enzymes there are sub-picosecond motions, on the same timescale as barrier passage, the couple directly to chemical transformation, and are thus part of the reaction coordinate. Approaches such as transition path sampling and committor analysis have greatly enhanced our understanding of these processes.
Figures
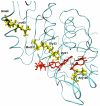
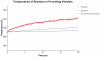
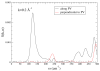
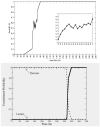
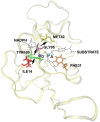

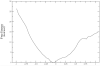
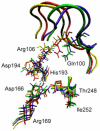
References
-
- Austin RH, Beeson KW, Eisenstein L, Frauenfe1der H, Gunsalus IC. Biochemistry. 1975;14:5355. - PubMed
-
- Agmon N, Hopfield J. J. Chern. Phys. 1983;79:2042.
-
- Brooks B, Bruccoleri R, Olafson B, States D, Swaminathan S, Karplus M. J. Comp. Chem. 1983;4:187–217.
-
- Gao J, Amara P, Alhambra C, Field M. J. Phys. Chem. A. 1998;102:4714.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

