Curcumin modulates α-synuclein aggregation and toxicity
- PMID: 23509976
- PMCID: PMC3605819
- DOI: 10.1021/cn3001203
Curcumin modulates α-synuclein aggregation and toxicity
Abstract
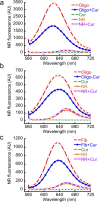
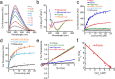
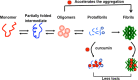
In human beings, Parkinson's disease (PD) is associated with the oligomerization and amyloid formation of α-synuclein (α-Syn). The polyphenolic Asian food ingredient curcumin has proven to be effective against a wide range of human diseases including cancers and neurological disorders. While curcumin has been shown to significantly reduce cell toxicity of α-Syn aggregates, its mechanism of action remains unexplored. Here, using a series of biophysical techniques, we demonstrate that curcumin reduces toxicity by binding to preformed oligomers and fibrils and altering their hydrophobic surface exposure. Further, our fluorescence and two-dimensional nuclear magnetic resonance (2D-NMR) data indicate that curcumin does not bind to monomeric α-Syn but binds specifically to oligomeric intermediates. The degree of curcumin binding correlates with the extent of α-Syn oligomerization, suggesting that the ordered structure of protein is required for effective curcumin binding. The acceleration of aggregation by curcumin may decrease the population of toxic oligomeric intermediates of α-Syn. Collectively; our results suggest that curcumin and related polyphenolic compounds can be pursued as candidate drug targets for treatment of PD and other neurological diseases.
Figures





References
-
- Cookson M. R. (2005) The biochemistry of Parkinson’s disease. Annu. Rev. Biochem. 74, 29–52. - PubMed
-
- Lansbury P. T.; Brice A. (2002) Genetics of Parkinson’s disease and biochemical studies of implicated gene products - Commentary. Curr. Opin Cell. Biol. 14, 653–660. - PubMed
-
- Feany M. B. (2000) Studying human neurodegenerative diseases in flies and worms. J. Neuropathol. Exp. Neurol. 59, 847–856. - PubMed
-
- Conway K. A.; Harper J. D.; Lansbury P. T. (1998) Accelerated in vitro fibril formation by a mutant α-synuclein linked to early-onset Parkinson disease. Nat. Med. 4, 1318–1320. - PubMed
-
- Conway K. A.; Lee S. J.; Rochet J. C.; Ding T. T.; Williamson R. E.; Lansbury P. T. (2000) Acceleration of oligomerization, not fibrillization, is a shared property of both a-synuclein mutations linked to early-onset Parkinson’s disease: Implications for pathogenesis and therapy. Proc. Natl. Acad. Sci. U.S.A. 97, 571–576. - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

