Quantum dot imaging platform for single-cell molecular profiling
- PMID: 23511483
- PMCID: PMC3615486
- DOI: 10.1038/ncomms2635
Quantum dot imaging platform for single-cell molecular profiling
Abstract
Study of normal cell physiology and disease pathogenesis heavily relies on untangling the complexity of intracellular molecular mechanisms and pathways. To achieve this goal, comprehensive molecular profiling of individual cells within the context of microenvironment is required. Here we report the development of a multicolour multicycle in situ imaging technology capable of creating detailed quantitative molecular profiles for individual cells at the resolution of optical imaging. A library of stoichiometric fluorescent probes is prepared by linking target-specific antibodies to a universal quantum dot-based platform via protein A in a quick and simple procedure. Surprisingly, despite the potential for multivalent binding between protein A and antibody and the intermediate affinity of this non-covalent bond, fully assembled probes do not aggregate or exchange antibodies, facilitating highly multiplexed parallel staining. This single-cell molecular profiling technology is expected to open new opportunities in systems biology, gene expression studies, signalling pathway analysis and molecular diagnostics.
Figures



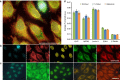



References
-
- de Souza N. Single-cell methods. Nat. Methods 9, 35 (2012).
-
- Liotta L. & Petricoin E. Molecular profiling of human cancer. Nat. Rev. Genet. 1, 48–56 (2000). - PubMed
-
- Zhang J., Campbell R. E., Ting A. Y. & Tsien R. Y. Creating new fluorescent probes for cell biology. Nat. Rev. Mol. Cell Biol. 3, 906–918 (2002). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

