Confirming genes influencing risk to cleft lip with/without cleft palate in a case-parent trio study
- PMID: 23512105
- PMCID: PMC3707506
- DOI: 10.1007/s00439-013-1283-6
Confirming genes influencing risk to cleft lip with/without cleft palate in a case-parent trio study
Abstract
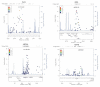
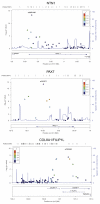
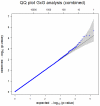
A collection of 1,108 case-parent trios ascertained through an isolated, nonsyndromic cleft lip with or without cleft palate (CL/P) was used to replicate the findings from a genome-wide association study (GWAS) conducted by Beaty et al. (Nat Genet 42:525-529, 2010), where four different genes/regions were identified as influencing risk to CL/P. Tagging SNPs for 33 different genes were genotyped (1,269 SNPs). All four of the genes originally identified as showing genome-wide significance (IRF6, ABCA4 and MAF, plus the 8q24 region) were confirmed in this independent sample of trios (who were primarily of European and Southeast Asian ancestry). In addition, eight genes classified as 'second tier' hits in the original study (PAX7, THADA, COL8A1/FILIP1L, DCAF4L2, GADD45G, NTN1, RBFOX3 and FOXE1) showed evidence of linkage and association in this replication sample. Meta-analysis between the original GWAS trios and these replication trios showed PAX7, COL8A1/FILIP1L and NTN1 achieved genome-wide significance. Tests for gene-environment interaction between these 33 genes and maternal smoking found evidence for interaction with two additional genes: GRID2 and ELAVL2 among European mothers (who had a higher rate of smoking than Asian mothers). Formal tests for gene-gene interaction (epistasis) failed to show evidence of statistical interaction in any simple fashion. This study confirms that many different genes influence risk to CL/P.
Figures


Similar articles
-
A multi-ethnic genome-wide association study identifies novel loci for non-syndromic cleft lip with or without cleft palate on 2p24.2, 17q23 and 19q13.Hum Mol Genet. 2016 Jul 1;25(13):2862-2872. doi: 10.1093/hmg/ddw104. Epub 2016 Mar 30. Hum Mol Genet. 2016. PMID: 27033726 Free PMC article.
-
A genome-wide association study of cleft lip with and without cleft palate identifies risk variants near MAFB and ABCA4.Nat Genet. 2010 Jun;42(6):525-9. doi: 10.1038/ng.580. Epub 2010 May 2. Nat Genet. 2010. PMID: 20436469 Free PMC article.
-
Gene-Gene Interaction Among WNT Genes for Oral Cleft in Trios.Genet Epidemiol. 2015 Jul;39(5):385-94. doi: 10.1002/gepi.21888. Epub 2015 Feb 6. Genet Epidemiol. 2015. PMID: 25663376 Free PMC article.
-
Evidence for SNP-SNP interaction identified through targeted sequencing of cleft case-parent trios.Genet Epidemiol. 2017 Apr;41(3):244-250. doi: 10.1002/gepi.22023. Epub 2016 Dec 26. Genet Epidemiol. 2017. PMID: 28019042 Free PMC article.
-
Joint testing of genotypic and gene-environment interaction identified novel association for BMP4 with non-syndromic CL/P in an Asian population using data from an International Cleft Consortium.PLoS One. 2014 Oct 10;9(10):e109038. doi: 10.1371/journal.pone.0109038. eCollection 2014. PLoS One. 2014. PMID: 25303326 Free PMC article.
Cited by
-
A genome-wide study of inherited deletions identified two regions associated with nonsyndromic isolated oral clefts.Birth Defects Res A Clin Mol Teratol. 2015 Apr;103(4):276-83. doi: 10.1002/bdra.23362. Epub 2015 Mar 16. Birth Defects Res A Clin Mol Teratol. 2015. PMID: 25776870 Free PMC article.
-
Genetics and signaling mechanisms of orofacial clefts.Birth Defects Res. 2020 Nov;112(19):1588-1634. doi: 10.1002/bdr2.1754. Epub 2020 Jul 15. Birth Defects Res. 2020. PMID: 32666711 Free PMC article. Review.
-
Facial Genetics: A Brief Overview.Front Genet. 2018 Oct 16;9:462. doi: 10.3389/fgene.2018.00462. eCollection 2018. Front Genet. 2018. PMID: 30386375 Free PMC article. Review.
-
Identification of Novel Genomic Variations in Susceptibility to Nonsyndromic Cleft Lip and Palate Patients.Pediatr Rep. 2021 Dec 8;13(4):650-657. doi: 10.3390/pediatric13040077. Pediatr Rep. 2021. PMID: 34941638 Free PMC article.
-
Association of Transforming Growth Factor Alpha Polymorphisms with Nonsyndromic Cleft Lip and Palate in Iranian Population.Avicenna J Med Biotechnol. 2015 Oct-Dec;7(4):168-72. Avicenna J Med Biotechnol. 2015. PMID: 26605011 Free PMC article.
References
-
- Beaty TH, Murray JC, Marazita ML, Munger RG, Ruczinski I, Hetmanski JB, Liang KY, Wu T, Murray T, Fallin MD, Redett RA, Raymond G, Schwender H, Jin SC, Rose M, Cooper ME, Dunnwald M, Mansilla MA, Leslie E, Bullard S, Lidral A, Moreno LM, Menezes R, Vieira AR, Petrin A, Wilcox A, Lie RT, Jabs EW, Wu-Chou YH, Wang H, Ye X, Huang S, Yeow V, Chong SS, Jee SH, Shi B, Christensen K, Doheny K, Pugh EW, Ling H, Castilla EE, Czeizel AE, Ma L, Field LL, Brody LC, Pangilinan F, Mills JL, Molloy AM, Kirke PN, Scott JM, Arcos-Burgos M, Scott AF. A genome wide association study of cleft lip with and without cleft palate identifies risk variants near MAFB and ABCA4. Nature Genetics. 2010;42:525–529. doi:10.38/ng.580. - PMC - PubMed
-
- Birnbaum S, Ludwig KU, Reutter H, Herms S, Steffens M, Rubini M, Baluardo C, Ferrian M, Almeida de Assis N, Alblas MA, Barth S, Freudenberg J, Lauster C, Schmidt G, Scheer M, Braumann B, Bergé SJ, Reich RH, Schiefke F, Hemprich A, Pötzsch S, Steegers-Theunissen RP, Pötzsch B, Moebus S, Horsthemke B, Kramer FJ, Wienker TF, Mossey PA, Propping P, Cichon S, Hoffmann P, Knapp M, Nöthen MM, Mangold E. Key susceptibility locus for nonsyndromic cleft lip with or without cleft palate on chromosome 8q24. Nature Genetics. 2009;41:473–477. doi:10.1038/ng.333. - PubMed
-
- Cordell HJ. Epistasis: What it means, what it doesn’t mean, and statistical methods to detect it in humans. Human Molecular Genetics. 2002;11:2463–2468. doi:10.1093/hmg/11.20.2463. - PubMed
Publication types
MeSH terms
Grants and funding
- R37 DE008559/DE/NIDCR NIH HHS/United States
- U01-DE-018993/DE/NIDCR NIH HHS/United States
- P30 ES005605/ES/NIEHS NIH HHS/United States
- U01 DE018993/DE/NIDCR NIH HHS/United States
- R01 DE014581/DE/NIDCR NIH HHS/United States
- R01-DE-016148/DE/NIDCR NIH HHS/United States
- R01 DE008559/DE/NIDCR NIH HHS/United States
- R01-DE-014581/DE/NIDCR NIH HHS/United States
- HHSN268200782096C/HG/NHGRI NIH HHS/United States
- R01 DE016148/DE/NIDCR NIH HHS/United States
- P50 DE016215/DE/NIDCR NIH HHS/United States
- P60 DE013076/DE/NIDCR NIH HHS/United States
- P50-DE016215/DE/NIDCR NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous

