Elevated rate of fixation of endogenous retroviral elements in Haplorhini TRIM5 and TRIM22 genomic sequences: impact on transcriptional regulation
- PMID: 23516500
- PMCID: PMC3597737
- DOI: 10.1371/journal.pone.0058532
Elevated rate of fixation of endogenous retroviral elements in Haplorhini TRIM5 and TRIM22 genomic sequences: impact on transcriptional regulation
Abstract
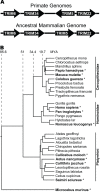
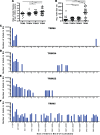
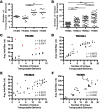
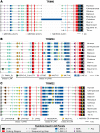
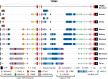
All genes in the TRIM6/TRIM34/TRIM5/TRIM22 locus are type I interferon inducible, with TRIM5 and TRIM22 possessing antiviral properties. Evolutionary studies involving the TRIM6/34/5/22 locus have predominantly focused on the coding sequence of the genes, finding that TRIM5 and TRIM22 have undergone high rates of both non-synonymous nucleotide replacements and in-frame insertions and deletions. We sought to understand if divergent evolutionary pressures on TRIM6/34/5/22 coding regions have selected for modifications in the non-coding regions of these genes and explore whether such non-coding changes may influence the biological function of these genes. The transcribed genomic regions, including the introns, of TRIM6, TRIM34, TRIM5, and TRIM22 from ten Haplorhini primates and one prosimian species were analyzed for transposable element content. In Haplorhini species, TRIM5 displayed an exaggerated interspecies variability, predominantly resulting from changes in the composition of transposable elements in the large first and fourth introns. Multiple lineage-specific endogenous retroviral long terminal repeats (LTRs) were identified in the first intron of TRIM5 and TRIM22. In the prosimian genome, we identified a duplication of TRIM5 with a concomitant loss of TRIM22. The transposable element content of the prosimian TRIM5 genes appears to largely represent the shared Haplorhini/prosimian ancestral state for this gene. Furthermore, we demonstrated that one such differentially fixed LTR provides for species-specific transcriptional regulation of TRIM22 in response to p53 activation. Our results identify a previously unrecognized source of species-specific variation in the antiviral TRIM genes, which can lead to alterations in their transcriptional regulation. These observations suggest that there has existed long-term pressure for exaptation of retroviral LTRs in the non-coding regions of these genes. This likely resulted from serial viral challenges and provided a mechanism for rapid alteration of transcriptional regulation. To our knowledge, this represents the first report of persistent evolutionary pressure for the capture of retroviral LTR insertions.
Conflict of interest statement
Figures



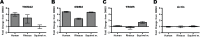
References
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

