Molecular dynamics simulations of DPPC bilayers using "LIME", a new coarse-grained model
- PMID: 23521567
- PMCID: PMC3703713
- DOI: 10.1021/jp309712b
Molecular dynamics simulations of DPPC bilayers using "LIME", a new coarse-grained model
Abstract
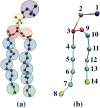
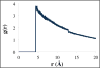
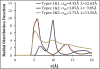
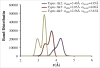
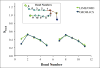
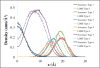
A new intermediate resolution model for phospholipids, LIME, designed for use with discontinuous molecular dynamics (DMD) simulations is presented. The implicit-solvent model was developed using a multiscale modeling approach in which the geometric and energetic parameters are obtained by collecting data from atomistic simulations of a system composed of 1,2-dipalmitoyl-sn-glycero-3-phosphocholine (DPPC) molecules and explicit water. In the model, 14 coarse-grained sites that are classified as 1 of 6 types represent DPPC. DMD simulations performed on a random solution of DPPC resulted in the formation of a defect-free bilayer in less than 4 h. The bilayer formed quantitatively reproduces the main structural properties (e.g., area per lipid, bilayer thickness, bond order parameters) that are observed experimentally. In addition, the bilayer transitions from a liquid-crystalline phase to a tilted gel phase when the temperature is reduced. Transbilayer movement of a lipid from the bottom leaflet to the top leaflet is observed when the temperature is increased.
Figures





Similar articles
-
Phase separation behavior of mixed lipid systems at neutral and low pH: coarse-grained simulations with DMD/LIME.Langmuir. 2015 Jan 27;31(3):1086-94. doi: 10.1021/la504082x. Epub 2015 Jan 15. Langmuir. 2015. PMID: 25549801 Free PMC article.
-
Surface tension effects on the phase transition of a DPPC bilayer with and without protein: a molecular dynamics simulation.Phys Chem Chem Phys. 2014 May 14;16(18):8434-40. doi: 10.1039/c3cp55524k. Phys Chem Chem Phys. 2014. PMID: 24668218
-
Comparing an All-Atom and a Coarse-Grained Description of Lipid Bilayers in Terms of Enthalpies and Entropies: From MD Simulations to 2D Lattice Models.J Chem Theory Comput. 2019 Nov 12;15(11):6393-6402. doi: 10.1021/acs.jctc.9b00390. Epub 2019 Oct 24. J Chem Theory Comput. 2019. PMID: 31593631
-
Atomistic Monte Carlo simulation of lipid membranes.Int J Mol Sci. 2014 Jan 24;15(2):1767-803. doi: 10.3390/ijms15021767. Int J Mol Sci. 2014. PMID: 24469314 Free PMC article. Review.
-
An NMR database for simulations of membrane dynamics.Biochim Biophys Acta. 2011 Mar;1808(3):818-39. doi: 10.1016/j.bbamem.2010.11.027. Epub 2010 Dec 4. Biochim Biophys Acta. 2011. PMID: 21134351 Free PMC article. Review.
Cited by
-
Development of a coarse-grained lipid model, LIME 2.0, for DSPE using multistate iterative Boltzmann inversion and discontinuous molecular dynamics simulations.Fluid Phase Equilib. 2020 Oct 15;521:112704. doi: 10.1016/j.fluid.2020.112704. Epub 2020 Jun 13. Fluid Phase Equilib. 2020. PMID: 37982069 Free PMC article.
-
Glutamate Permeability of Chicken Best1.Exp Neurobiol. 2022 Oct 31;31(5):277-288. doi: 10.5607/en22038. Exp Neurobiol. 2022. PMID: 36351838 Free PMC article.
-
Modelling lipid systems in fluid with Lattice Boltzmann Molecular Dynamics simulations and hydrodynamics.Sci Rep. 2019 Nov 11;9(1):16450. doi: 10.1038/s41598-019-52760-y. Sci Rep. 2019. PMID: 31712588 Free PMC article.
-
Phase separation behavior of mixed lipid systems at neutral and low pH: coarse-grained simulations with DMD/LIME.Langmuir. 2015 Jan 27;31(3):1086-94. doi: 10.1021/la504082x. Epub 2015 Jan 15. Langmuir. 2015. PMID: 25549801 Free PMC article.
-
Transferring the PRIMO Coarse-Grained Force Field to the Membrane Environment: Simulations of Membrane Proteins and Helix-Helix Association.J Chem Theory Comput. 2014 Aug 12;10(8):3459-3472. doi: 10.1021/ct500443v. Epub 2014 Jun 16. J Chem Theory Comput. 2014. PMID: 25136271 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

