Global analysis of phosphorylation networks in humans
- PMID: 23524292
- PMCID: PMC3815481
- DOI: 10.1016/j.bbapap.2013.03.009
Global analysis of phosphorylation networks in humans
Abstract
Phosphorylation-mediated signaling plays a crucial role in nearly every aspect of cellular physiology. A recent study based on protein microarray experiments identified a large number of kinase-substrate relationships (KSRs), and built a comprehensive and reliable phosphorylation network in humans. Analysis of this network, in conjunction with additional resources, revealed several key features. First, comparison of the human and yeast phosphorylation networks uncovered an evolutionarily conserved signaling backbone dominated by kinase-to-kinase relationships. Second, although most of the KSRs themselves are not conserved, the functions enriched in the substrates for a given kinase are often conserved. Third, the prevalence of kinase-transcription factor regulatory modules suggests that phosphorylation and transcriptional regulatory networks are inherently wired together to form integrated regulatory circuits. Overall, the phosphorylation networks described in this work promise to offer new insights into the properties of kinase signaling pathways, at both the global and the protein levels. This article is part of a Special Issue entitled: Computational Proteomics, Systems Biology & Clinical Implications. Guest Editor: Yudong Cai.
Keywords: Conservation; Kinase–substrate relationships (KSRs); Network module; Phosphorylation network.
Copyright © 2013 Elsevier B.V. All rights reserved.
Figures
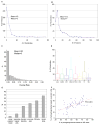





References
-
- Mullor JL, Sanchez P, Ruiz i Altaba A. Pathways and consequences: Hedgehog signaling in human disease. Trends Cell Biol. 2002;12:562–569. - PubMed
-
- Nusse R. Wnt signaling in disease and in development. Cell Res. 2005;15:28–32. - PubMed
-
- Smahi A, Courtois G, Rabia SH, Doffinger R, Bodemer C, Munnich A, Casanova JL, Israel A. The NF-kappaB signalling pathway in human diseases: from incontinentia pigmenti to ectodermal dysplasias and immune-deficiency syndromes. Hum Mol Genet. 2002;11:2371–2375. - PubMed
-
- Tsatsanis C, Spandidos DA. The role of oncogenic kinases in human cancer (Review) Int J Mol Med. 2000;5:583–590. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

