Early gene expression divergence between allopatric populations of the house mouse (Mus musculus domesticus)
- PMID: 23532401
- PMCID: PMC3605846
- DOI: 10.1002/ece3.447
Early gene expression divergence between allopatric populations of the house mouse (Mus musculus domesticus)
Abstract
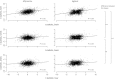
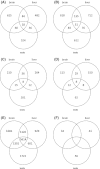
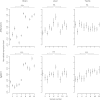
Divergence of gene expression is known to contribute to the differentiation and separation of populations and species, although the dynamics of this process in early stages of population divergence remains unclear. We analyzed gene expression differences in three organs (brain, liver, and testis) between two natural populations of Mus musculus domesticus that have been separated for at most 3000 years. We used two different microarray platforms to corroborate the results at a large scale and identified hundreds of genes with significant expression differences between the populations. We find that although the three tissues have similar number of differentially expressed genes, brain and liver have more tissue-specific genes than testis. Most genes show changes in a single tissue only, even when expressed in all tissues, supporting the notion that tissue-specific enhancers act as separable targets of evolution. In terms of functional categories, in brain and to a smaller extent in liver, we find transcription factors and their targets to be particularly variable between populations, similar to previous findings in primates. Testis, however, has a different set of differently expressed genes, both with respect to functional categories and overall correlation with the other tissues, the latter indicating that gene expression divergence of potential importance might be present in other datasets where no differences in fraction of differentially expressed genes were reported. Our results show that a significant amount of gene expression divergence quickly accumulates between allopatric populations.
Keywords: Evolution; gene expression; population divergence; wild mice.
Figures


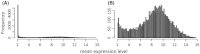
References
-
- Boursot P, Auffray J, Britton-Davidian J, Bonhomme F. The evolution of house mice. Ann. Rev. Ecol. Evol. Syst. 1993;24:119–152.
-
- Brawand D, Soumillon M, Necsulea A, Julien P, Csárdi G, Harrigan P, et al. The evolution of gene expression levels in mammalian organs. Nature. 2011;478:343–348. - PubMed
-
- Carroll SB. Evo-devo and an expanding evolutionary synthesis: a genetic theory of morphological evolution. Cell. 2008;134:25–36. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

