A novel automethylation reaction in the Aspergillus nidulans LaeA protein generates S-methylmethionine
- PMID: 23532849
- PMCID: PMC3656261
- DOI: 10.1074/jbc.M113.465765
A novel automethylation reaction in the Aspergillus nidulans LaeA protein generates S-methylmethionine
Abstract
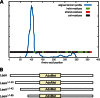
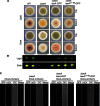
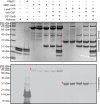
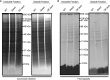
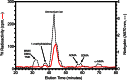
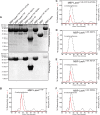
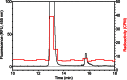
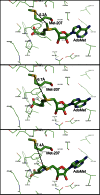
The filamentous fungi in the genus Aspergillus are opportunistic plant and animal pathogens that can adapt to their environment by producing various secondary metabolites, including lovastatin, penicillin, and aflatoxin. The synthesis of these small molecules is dependent on gene clusters that are globally regulated by the LaeA protein. Null mutants of LaeA in all pathogenic fungi examined to date show decreased virulence coupled with reduced secondary metabolism. Although the amino acid sequence of LaeA contains the motifs characteristic of seven-β-strand methyltransferases, a methyl-accepting substrate of LaeA has not been identified. In this work we did not find a methyl-accepting substrate in Aspergillus nidulans with various assays, including in vivo S-adenosyl-[methyl-(3)H]methionine labeling, targeted in vitro methylation experiments using putative protein substrates, or in vitro methylation assays using whole cell extracts grown under different conditions. However, in each experiment LaeA was shown to self-methylate. Amino acid hydrolysis of radioactively labeled LaeA followed by cation exchange and reverse phase chromatography identified methionine as the modified residue. Point mutations show that the major site of modification of LaeA is on methionine 207. However, in vivo complementation showed that methionine 207 is not required for the biological function of LaeA. LaeA is the first protein to exhibit automethylation at a methionine residue. These findings not only indicate LaeA may perform novel chemistry with S-adenosylmethionine but also provide new insights into the physiological function of LaeA.
Keywords: Aspergillus; Automethylation; Gene Regulation; Histone Methylation; LaeA; Methionine; Protein Methylation; S-Adenosylmethionine; S-Methylmethionine; Secondary Metabolism.
Figures




Similar articles
-
LaeA Controls Citric Acid Production through Regulation of the Citrate Exporter-Encoding cexA Gene in Aspergillus luchuensis mut. kawachii.Appl Environ Microbiol. 2020 Feb 18;86(5):e01950-19. doi: 10.1128/AEM.01950-19. Print 2020 Feb 18. Appl Environ Microbiol. 2020. PMID: 31862728 Free PMC article.
-
Functional analyses of Trichoderma reesei LAE1 reveal conserved and contrasting roles of this regulator.G3 (Bethesda). 2013 Feb;3(2):369-78. doi: 10.1534/g3.112.005140. Epub 2013 Feb 1. G3 (Bethesda). 2013. PMID: 23390613 Free PMC article.
-
Accurate prediction of the Aspergillus nidulans terrequinone gene cluster boundaries using the transcriptional regulator LaeA.Fungal Genet Biol. 2007 Nov;44(11):1134-45. doi: 10.1016/j.fgb.2006.12.010. Epub 2007 Jan 8. Fungal Genet Biol. 2007. PMID: 17291795
-
Key role of LaeA and velvet complex proteins on expression of β-lactam and PR-toxin genes in Penicillium chrysogenum: cross-talk regulation of secondary metabolite pathways.J Ind Microbiol Biotechnol. 2017 May;44(4-5):525-535. doi: 10.1007/s10295-016-1830-y. Epub 2016 Aug 26. J Ind Microbiol Biotechnol. 2017. PMID: 27565675 Review.
-
Beyond morphogenesis and secondary metabolism: function of Velvet proteins and LaeA in fungal pathogenesis.Appl Environ Microbiol. 2024 Oct 23;90(10):e0081924. doi: 10.1128/aem.00819-24. Epub 2024 Sep 4. Appl Environ Microbiol. 2024. PMID: 39230285 Free PMC article. Review.
Cited by
-
The Role of Chromatin and Transcriptional Control in the Formation of Sexual Fruiting Bodies in Fungi.Microbiol Mol Biol Rev. 2022 Dec 21;86(4):e0010422. doi: 10.1128/mmbr.00104-22. Epub 2022 Nov 21. Microbiol Mol Biol Rev. 2022. PMID: 36409109 Free PMC article. Review.
-
The BcLAE1 is involved in the regulation of ABA biosynthesis in Botrytis cinerea TB-31.Front Microbiol. 2022 Aug 4;13:969499. doi: 10.3389/fmicb.2022.969499. eCollection 2022. Front Microbiol. 2022. PMID: 35992717 Free PMC article.
-
The putative methyltransferase LaeA regulates mycelium growth and cellulase production in Myceliophthora thermophila.Biotechnol Biofuels Bioprod. 2023 Apr 3;16(1):58. doi: 10.1186/s13068-023-02313-3. Biotechnol Biofuels Bioprod. 2023. PMID: 37013645 Free PMC article.
-
The VELVET A orthologue VEL1 of Trichoderma reesei regulates fungal development and is essential for cellulase gene expression.PLoS One. 2014 Nov 11;9(11):e112799. doi: 10.1371/journal.pone.0112799. eCollection 2014. PLoS One. 2014. PMID: 25386652 Free PMC article.
-
LaeA Controls Citric Acid Production through Regulation of the Citrate Exporter-Encoding cexA Gene in Aspergillus luchuensis mut. kawachii.Appl Environ Microbiol. 2020 Feb 18;86(5):e01950-19. doi: 10.1128/AEM.01950-19. Print 2020 Feb 18. Appl Environ Microbiol. 2020. PMID: 31862728 Free PMC article.
References
-
- Latgé J. P. (2001) The pathobiology of Aspergillus fumigatus. Trends Microbiol. 9, 382–389 - PubMed
-
- Alberts A. W., Chen J., Kuron G., Hunt V., Huff J., Hoffman C., Rothrock J., Lopez M., Joshua H., Harris E., Patchett A., Monaghan R., Currie S., Stapley E., Albers-Schonberg G., Hensens O., Hirshfield J., Hoogsteen K., Liesch J., Springer J. (1980) Mevinolin: a highly potent competitive inhibitor of hydroxymethylglutaryl-coenzyme A reductase and a cholesterol-lowering agent. Proc. Natl. Acad. Sci. U.S.A. 77, 3957–3961 - PMC - PubMed
-
- Keller N. P., Turner G., Bennett J. W. (2005) Fungal secondary metabolism–from biochemistry to genomics. Nat. Rev. Microbiol. 3, 937–947 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

