Structure and RNA-binding properties of the bacterial LSm protein Hfq
- PMID: 23535768
- PMCID: PMC3710368
- DOI: 10.4161/rna.24201
Structure and RNA-binding properties of the bacterial LSm protein Hfq
Abstract
Over the past years, small non-coding RNAs (sRNAs) emerged as important modulators of gene expression in bacteria. Guided by partial sequence complementarity, these sRNAs interact with target mRNAs and eventually affect transcript stability and translation. The physiological function of sRNAs depends on the protein Hfq, which binds sRNAs in the cell and promotes the interaction with their mRNA targets. This important physiological function of Hfq as a central hub of sRNA-mediated regulation made it one of the most intensely studied proteins in bacteria. Recently, a new model for sRNA binding by Hfq has been proposed that involves the direct recognition of the sRNA 3' end and interactions of the sRNA body with the lateral RNA-binding surface of Hfq. This review summarizes the current understanding of the RNA binding properties of Hfq and its (s)RNA complexes. Moreover, the implications of the new binding model for sRNA-mediated regulation are discussed.
Keywords: 3′ end recognition; LSm ring; RNA chaperone; RNA degradation; crystal structure; gene regulation; non-coding RNAs; prokaryotes.
Figures
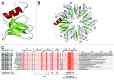

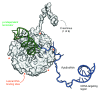
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
