Proteasome-mediated proteolysis of SRSF5 splicing factor intriguingly co-occurs with SRSF5 mRNA upregulation during late erythroid differentiation
- PMID: 23536862
- PMCID: PMC3594168
- DOI: 10.1371/journal.pone.0059137
Proteasome-mediated proteolysis of SRSF5 splicing factor intriguingly co-occurs with SRSF5 mRNA upregulation during late erythroid differentiation
Abstract
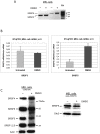
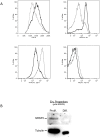
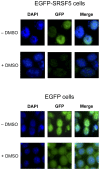
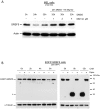
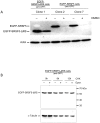
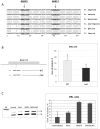
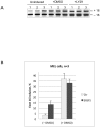
SR proteins exhibit diverse functions ranging from their role in constitutive and alternative splicing, to virtually all aspects of mRNA metabolism. These findings have attracted growing interest in deciphering the regulatory mechanisms that control the tissue-specific expression of these SR proteins. In this study, we show that SRSF5 protein decreases drastically during erythroid cell differentiation, contrasting with a concomitant upregulation of SRSF5 mRNA level. Proteasome chemical inhibition provided strong evidence that endogenous SRSF5 protein, as well as protein deriving from stably transfected SRSF5 cDNA, are both targeted to proteolysis as the cells undergo terminal differentiation. Consistently, functional experiments show that overexpression of SRSF5 enhances a specific endogenous pre-mRNA splicing event in proliferating cells, but not in differentiating cells, due to proteasome-mediated targeting of both endogenous and transfection-derived SRSF5. Further investigation of the relationship between SRSF5 structure and its post-translation regulation and function, suggested that the RNA recognition motifs of SRSF5 are sufficient to activate pre-mRNA splicing, whereas proteasome-mediated proteolysis of SRSF5 requires the presence of the C-terminal RS domain of the protein. Phosphorylation of SR proteins is a key post-translation regulation that promotes their activity and subcellular availability. We here show that inhibition of the CDC2-like kinase (CLK) family and mutation of the AKT phosphorylation site Ser86 on SRSF5, have no effect on SRSF5 stability. We reasoned that at least AKT and CLK signaling pathways are not involved in proteasome-induced turnover of SRSF5 during late erythroid development.
Conflict of interest statement
Figures


References
-
- Lin S, Fu XD (2007) SR proteins and related factors in alternative splicing. Adv Exp Med Biol 623: 107–122. - PubMed
-
- Yoshida K, Sanada M, Shiraishi Y, Nowak D, Nagata Y, et al. (2011) Frequent pathway mutations of splicing machinery in myelodysplasia. Nature 478: 64–69. - PubMed
-
- Long JC, Caceres JF (2009) The SR protein family of splicing factors: master regulators of gene expression. Biochem J 417: 15–27. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

