Bacterial protein-O-mannosylating enzyme is crucial for virulence of Mycobacterium tuberculosis
- PMID: 23550160
- PMCID: PMC3631654
- DOI: 10.1073/pnas.1219704110
Bacterial protein-O-mannosylating enzyme is crucial for virulence of Mycobacterium tuberculosis
Abstract
A posttranslational protein O-mannosylation process resembling that found in fungi and animals has been reported in the major human pathogen Mycobacterium tuberculosis (Mtb) and related actinobacteria. However, the role and incidence of this process, which is essential in eukaryotes, have never been explored in Mtb. We thus analyzed the impact of interrupting O-mannosylation in the nonpathogenic saprophyte Mycobacterium smegmatis and in the human pathogen Mtb by inactivating the respective putative protein mannosyl transferase genes Msmeg_5447 and Rv1002c. Loss of protein O-mannosylation in both mutant strains was unambiguously demonstrated by efficient mass spectrometry-based glycoproteomics analysis. Unexpectedly, although the M. smegmatis phenotype was unaffected by the lack of manno-proteins, the Mtb mutant had severely impacted growth in vitro and in cellulo associated with a strong attenuation of its pathogenicity in immunocompromised mice. These data are unique in providing evidence of the biological significance of protein O-mannosylation in mycobacteria and demonstrate the crucial contribution of this protein posttranslational modification to Mtb virulence in the host.
Conflict of interest statement
The authors declare no conflict of interest.
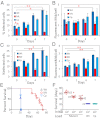
Figures



Similar articles
-
Development of a novel target-based cell assay, reporter of the activity of Mycobacterium tuberculosis protein-O-mannosyltransferase.Glycobiology. 2023 Dec 30;33(12):1139-1154. doi: 10.1093/glycob/cwad072. Glycobiology. 2023. PMID: 37698262
-
Export-mediated assembly of mycobacterial glycoproteins parallels eukaryotic pathways.Science. 2005 Aug 5;309(5736):941-3. doi: 10.1126/science.1114347. Science. 2005. PMID: 16081738
-
Potential Plasticity of the Mannoprotein Repertoire Associated to Mycobacterium tuberculosis Virulence Unveiled by Mass Spectrometry-Based Glycoproteomics.Molecules. 2020 May 18;25(10):2348. doi: 10.3390/molecules25102348. Molecules. 2020. PMID: 32443484 Free PMC article.
-
Protein O-mannosylation: conserved from bacteria to humans.Glycobiology. 2009 Aug;19(8):816-28. doi: 10.1093/glycob/cwp066. Epub 2009 May 9. Glycobiology. 2009. PMID: 19429925 Review.
-
Protein-O-mannosyltransferases in virulence and development.Cell Mol Life Sci. 2008 Feb;65(4):528-44. doi: 10.1007/s00018-007-7409-z. Cell Mol Life Sci. 2008. PMID: 17975704 Free PMC article. Review.
Cited by
-
Quantitative mass spectrometry reveals plasticity of metabolic networks in Mycobacterium smegmatis.Mol Cell Proteomics. 2014 Nov;13(11):3014-28. doi: 10.1074/mcp.M113.034082. Epub 2014 Jul 5. Mol Cell Proteomics. 2014. PMID: 24997995 Free PMC article.
-
Differential substrate preferences IN ACTINOBACTERIAL protein O-MANNOSYLTRANSFERASES and alteration of protein-O-MANNOSYLATION by choice of secretion pathway.Glycobiology. 2025 Jan 13;35(1):cwae095. doi: 10.1093/glycob/cwae095. Glycobiology. 2025. PMID: 39673494 Free PMC article.
-
Deciphering the molecular basis of mycobacteria and lipoglycan recognition by the C-type lectin Dectin-2.Sci Rep. 2018 Nov 15;8(1):16840. doi: 10.1038/s41598-018-35393-5. Sci Rep. 2018. PMID: 30443026 Free PMC article.
-
Effect of Protein O-Mannosyltransferase (MSMEG_5447) on M. smegmatis and Its Survival in Macrophages.Front Microbiol. 2021 Jun 30;12:657726. doi: 10.3389/fmicb.2021.657726. eCollection 2021. Front Microbiol. 2021. PMID: 34276591 Free PMC article.
-
Comparison of the transcriptome, lipidome, and c-di-GMP production between BCGΔBCG1419c and BCG, with Mincle- and Myd88-dependent induction of proinflammatory cytokines in murine macrophages.Sci Rep. 2024 May 24;14(1):11898. doi: 10.1038/s41598-024-61815-8. Sci Rep. 2024. PMID: 38789479 Free PMC article.
References
-
- Ragas A, Roussel L, Puzo G, Rivière M. The Mycobacterium tuberculosis cell-surface glycoprotein apa as a potential adhesin to colonize target cells via the innate immune system pulmonary C-type lectin surfactant protein A. J Biol Chem. 2007;282(8):5133–5142. - PubMed
-
- Nguyen L, Pieters J. The Trojan horse: Survival tactics of pathogenic mycobacteria in macrophages. Trends Cell Biol. 2005;15(5):269–276. - PubMed
-
- Espitia C, Servín-González L, Mancilla R. New insights into protein O-mannosylation in actinomycetes. Mol Biosyst. 2010;6(5):775–781. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

