Sumoylation of AMPKβ2 subunit enhances AMP-activated protein kinase activity
- PMID: 23552691
- PMCID: PMC3667731
- DOI: 10.1091/mbc.E12-11-0806
Sumoylation of AMPKβ2 subunit enhances AMP-activated protein kinase activity
Abstract
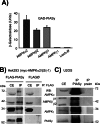
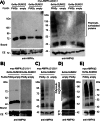
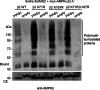
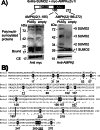
AMP-activated protein kinase (AMPK) is a sensor of cellular energy status. It is a heterotrimer composed of a catalytic α and two regulatory subunits (β and γ). AMPK activity is regulated allosterically by AMP and by the phosphorylation of residue Thr-172 within the catalytic domain of the AMPKα subunit by upstream kinases. We present evidence that the AMPKβ2 subunit may be posttranslationally modified by sumoylation. This process is carried out by the E3-small ubiquitin-like modifier (SUMO) ligase protein inhibitor of activated STAT PIASy, which modifies the AMPKβ2 subunit by the attachment of SUMO2 but not SUMO1 moieties. Of interest, AMPKβ1 is not a substrate for this modification. We also demonstrate that sumoylation of AMPKβ2 enhances the activity of the trimeric α2β2γ1 AMPK complex. In addition, our results indicate that sumoylation is antagonist and competes with the ubiquitination of the AMPKβ2 subunit. This adds a new layer of complexity to the regulation of the activity of the AMPK complex, since conditions that promote ubiquitination result in inactivation, whereas those that promote sumoylation result in the activation of the AMPK complex.
Figures




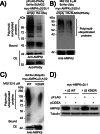
References
-
- Al-Hakim AK, Zagorska A, Chapman L, Deak M, Peggie M, Alessi DR. Control of AMPK-related kinases by USP9X and atypical Lys(29)/Lys(33)-linked polyubiquitin chains. Biochem J. 2008;411:249–260. - PubMed
-
- Bateman A. The structure of a domain common to archaebacteria and the homocystinuria disease protein. Trends Biochem Sci. 1997;22:12–13. - PubMed
-
- Bergink S, Jentsch S. Principles of ubiquitin and SUMO modifications in DNA repair. Nature. 2009;458:461–467. - PubMed
-
- Carling D, Thornton C, Woods A, Sanders MJ. AMP-activated protein kinase: new regulation, new roles. Biochem J. 2012;445:11–27. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

