Clustering and alignment of polymorphic sequences for HLA-DRB1 genotyping
- PMID: 23555798
- PMCID: PMC3610899
- DOI: 10.1371/journal.pone.0059835
Clustering and alignment of polymorphic sequences for HLA-DRB1 genotyping
Abstract
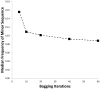
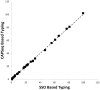
Located on Chromosome 6p21, classical human leukocyte antigen genes are highly polymorphic. HLA alleles associate with a variety of phenotypes, such as narcolepsy, autoimmunity, as well as immunologic response to infectious disease. Moreover, high resolution genotyping of these loci is critical to achieving long-term survival of allogeneic transplants. Development of methods to obtain high resolution analysis of HLA genotypes will lead to improved understanding of how select alleles contribute to human health and disease risk. Genomic DNAs were obtained from a cohort of n = 383 subjects recruited as part of an Ulcerative Colitis study and analyzed for HLA-DRB1. HLA genotypes were determined using sequence specific oligonucleotide probes and by next-generation sequencing using the Roche/454 GSFLX instrument. The Clustering and Alignment of Polymorphic Sequences (CAPSeq) software application was developed to analyze next-generation sequencing data. The application generates HLA sequence specific 6-digit genotype information from next-generation sequencing data using MUMmer to align sequences and the R package diffusionMap to classify sequences into their respective allelic groups. The incorporation of Bootstrap Aggregating, Bagging to aid in sorting of sequences into allele classes resulted in improved genotyping accuracy. Using Bagging iterations equal to 60, the genotyping results obtained using CAPSeq when compared with sequence specific oligonucleotide probe characterized 4-digit genotypes exhibited high rates of concordance, matching at 759 out of 766 (99.1%) alleles.
Conflict of interest statement
Figures

Similar articles
-
Concordance of next generation sequence-based and sequence specific oligonucleotide probe-based HLA-DRB1 genotyping.Hum Immunol. 2015 Dec;76(12):939-44. doi: 10.1016/j.humimm.2015.07.235. Epub 2015 Aug 4. Hum Immunol. 2015. PMID: 26247828 Free PMC article.
-
Reference Grade Characterization of Polymorphisms in Full-Length HLA Class I and II Genes With Short-Read Sequencing on the ION PGM System and Long-Reads Generated by Single Molecule, Real-Time Sequencing on the PacBio Platform.Front Immunol. 2018 Oct 4;9:2294. doi: 10.3389/fimmu.2018.02294. eCollection 2018. Front Immunol. 2018. PMID: 30337930 Free PMC article.
-
HLA-DRB1, -DRB3, -DRB4 and -DRB5 genotyping at a super-high resolution level by long range PCR and high-throughput sequencing.Tissue Antigens. 2014 Jan;83(1):10-6. doi: 10.1111/tan.12258. Epub 2013 Nov 30. Tissue Antigens. 2014. PMID: 24355003
-
High-resolution, high-throughput HLA genotyping by next-generation sequencing.Tissue Antigens. 2009 Nov;74(5):393-403. doi: 10.1111/j.1399-0039.2009.01345.x. Tissue Antigens. 2009. PMID: 19845894 Free PMC article.
-
HLA-DRB1 alleles may influence disease phenotype in patients with inflammatory bowel disease: a critical reappraisal with review of the literature.Dis Colon Rectum. 2005 Jan;48(1):57-64; discussion 64-5. doi: 10.1007/s10350-004-0747-0. Dis Colon Rectum. 2005. PMID: 15690658 Review.
Cited by
-
The Immune Response and the Pathogenesis of Idiopathic Inflammatory Myositis: a Critical Review.Clin Rev Allergy Immunol. 2017 Feb;52(1):58-70. doi: 10.1007/s12016-016-8527-x. Clin Rev Allergy Immunol. 2017. PMID: 26780034 Review.
-
Sleep disorders in multiple sclerosis. Review.Curr Neurol Neurosci Rep. 2015 May;15(5):21. doi: 10.1007/s11910-015-0546-0. Curr Neurol Neurosci Rep. 2015. PMID: 25773000 Review.
-
Autoimmune/inflammatory syndrome induced by adjuvant (ASIA) evolution after silicone implants. Who is at risk?Clin Rheumatol. 2015 Oct;34(10):1661-6. doi: 10.1007/s10067-015-2931-0. Epub 2015 Apr 16. Clin Rheumatol. 2015. PMID: 25877803 Review.
-
The impact of next-generation sequencing technologies on HLA research.J Hum Genet. 2015 Nov;60(11):665-73. doi: 10.1038/jhg.2015.102. Epub 2015 Aug 27. J Hum Genet. 2015. PMID: 26311539 Free PMC article. Review.
References
-
- Rossini AA, Greiner DL, Mordes JP (1999) Induction of immunologic tolerance for transplantation. Physiol Rev 79: 99–141. - PubMed
-
- Shiina T, Hosomichi K, Inoko H, Kulski JK (2009) The HLA genomic loci map: expression, interaction, diversity and disease. J Hum Genet 54: 15–39. - PubMed
-
- Becquemont L (2010) HLA: a pharmacogenomics success story. Pharmacogenomics 11: 277–281. - PubMed
-
- Karanes C, Nelson GO, Chitphakdithai P, Agura E, Ballen KK, et al. (2008) Twenty years of unrelated donor hematopoietic cell transplantation for adult recipients facilitated by the National Marrow Donor Program. Biol Blood Marrow Transplant 14: 8–15. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials

