Characterisation of ATP-dependent Mur ligases involved in the biogenesis of cell wall peptidoglycan in Mycobacterium tuberculosis
- PMID: 23555903
- PMCID: PMC3605390
- DOI: 10.1371/journal.pone.0060143
Characterisation of ATP-dependent Mur ligases involved in the biogenesis of cell wall peptidoglycan in Mycobacterium tuberculosis
Erratum in
-
Correction: Characterisation of ATP-Dependent Mur Ligases Involved in the Biogenesis of Cell Wall Peptidoglycan in Mycobacterium tuberculosis.PLoS One. 2024 Mar 26;19(3):e0301375. doi: 10.1371/journal.pone.0301375. eCollection 2024. PLoS One. 2024. PMID: 38530839 Free PMC article.
Abstract
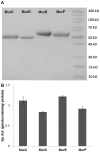
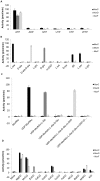
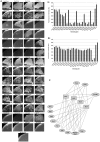
ATP-dependent Mur ligases (Mur synthetases) play essential roles in the biosynthesis of cell wall peptidoglycan (PG) as they catalyze the ligation of key amino acid residues to the stem peptide at the expense of ATP hydrolysis, thus representing potential targets for antibacterial drug discovery. In this study we characterized the division/cell wall (dcw) operon and identified a promoter driving the co-transcription of mur synthetases along with key cell division genes such as ftsQ and ftsW. Furthermore, we have extended our previous investigations of MurE to MurC, MurD and MurF synthetases from Mycobacterium tuberculosis. Functional analyses of the pure recombinant enzymes revealed that the presence of divalent cations is an absolute requirement for their activities. We also observed that higher concentrations of ATP and UDP-sugar substrates were inhibitory for the activities of all Mur synthetases suggesting stringent control of the cytoplasmic steps of the peptidoglycan biosynthetic pathway. In line with the previous findings on the regulation of mycobacterial MurD and corynebacterial MurC synthetases via phosphorylation, we found that all of the Mur synthetases interacted with the Ser/Thr protein kinases, PknA and PknB. In addition, we critically analyzed the interaction network of all of the Mur synthetases with proteins involved in cell division and cell wall PG biosynthesis to re-evaluate the importance of these key enzymes as novel therapeutic targets in anti-tubercular drug discovery.
Conflict of interest statement
Figures


References
-
- WHO (2012) Global tuberculosis control. WHO report 2012.
-
- Koul A, Arnoult E, Lounis N, Guillemont J, Andries K (2011) The challenge of new drug discovery for tuberculosis. Nature 469: 483–490. - PubMed
-
- Anishetty S, Pulimi M, Pennathur G (2005) Potential drug targets in Mycobacterium tuberculosis through metabolic pathway analysis. Comput Biol Chem 29: 368–378. - PubMed
-
- Guzman JD, Gupta A, Evangelopoulos D, Basavannacharya C, Pabon LC, et al. (2010) Anti-tubercular screening of natural products from Colombian plants: 3-methoxynordomesticine, an inhibitor of MurE ligase of Mycobacterium tuberculosis . J Antimicrob Chemother 65: 2101–2107. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

