Phosphorylation of mitophagy and pexophagy receptors coordinates their interaction with Atg8 and Atg11
- PMID: 23559066
- PMCID: PMC3642380
- DOI: 10.1038/embor.2013.40
Phosphorylation of mitophagy and pexophagy receptors coordinates their interaction with Atg8 and Atg11
Abstract
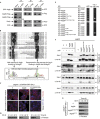
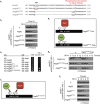
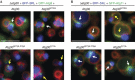
The selective autophagy receptors Atg19 and Atg32 interact with two proteins of the core autophagic machinery: the scaffold protein Atg11 and the ubiquitin-like protein Atg8. We found that the Pichia pastoris pexophagy receptor, Atg30, also interacts with Atg8. Both Atg30 and Atg32 interactions are regulated by phosphorylation close to Atg8-interaction motifs. Extending this finding to Saccharomyces cerevisiae, we confirmed phosphoregulation for the mitophagy and pexophagy receptors, Atg32 and Atg36. Each Atg30 molecule must interact with both Atg8 and Atg11 for full functionality, and these interactions occur independently and not simultaneously, but rather in random order. We present a common model for the phosphoregulation of selective autophagy receptors.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

