Bioactive conformations of two seminal delta opioid receptor penta-peptides inferred from free-energy profiles
- PMID: 23564013
- PMCID: PMC3812360
- DOI: 10.1002/bip.22251
Bioactive conformations of two seminal delta opioid receptor penta-peptides inferred from free-energy profiles
Abstract
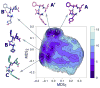
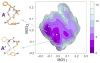
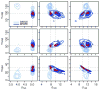
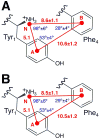
Delta-opioid (DOP) receptors are members of the G protein-coupled receptor (GPCR) sub-family of opioid receptors, and are evolutionarily related, with homology exceeding 70%, to cognate mu-opioid (MOP), kappa-opioid (KOP), and nociceptin opioid (NOP) receptors. DOP receptors are considered attractive drug targets for pain management because agonists at these receptors are reported to exhibit strong antinociceptive activity with relatively few side effects. Among the most potent analgesics targeting the DOP receptor are the linear and cyclic enkephalin analogs known as DADLE (Tyr-D-Ala-Gly-Phe-D-Leu) and DPDPE (Tyr-D-Pen-Gly-Phe-D-Pen), respectively. Several computational and experimental studies have been carried out over the years to characterize the conformational profile of these penta-peptides with the ultimate goal of designing potent peptidomimetic agonists for the DOP receptor. The computational studies published to date, however, have investigated only a limited range of timescales and used over-simplified representations of the solvent environment. We provide here a thorough exploration of the conformational space of DADLE and DPDPE in an explicit solvent, using microsecond-scale molecular dynamics and bias-exchange metadynamics simulations. Free-energy profiles derived from these simulations point to a small number of DADLE and DPDPE conformational minima in solution, which are separated by relatively small energy barriers. Candidate bioactive forms of these peptides are selected from identified common spatial arrangements of key pharmacophoric points within all sampled conformations.
Keywords: molecular dynamics; opioids; peptides.
Copyright © 2013 Wiley Periodicals, Inc.
Figures
References
-
- Porreca F, Bilsky EJ, Raffa RB, Lai J. The Pharmacology of Opioid Peptides. Harwood Academic Publishers; New Jersey: 1995.
-
- Hughes J, Smith TW, Kosterlitz HW, Fothergill LA, Morgan BA, Morris HR. Nature. 1975;258:577–580. - PubMed
-
- Graham WH, Carter ES, 2nd, Hicks RP. Biopolymers. 1992;32:1755–1764. - PubMed
-
- Naito A, Nishimura K. Curr Top Med Chem. 2004;4:135–145. - PubMed
-
- Deschamps JR, George C, Flippen-Anderson JL. Biopolymers. 1996;40:121–139. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

