Structure of the Legionella effector AnkX reveals the mechanism of phosphocholine transfer by the FIC domain
- PMID: 23572077
- PMCID: PMC3655470
- DOI: 10.1038/emboj.2013.82
Structure of the Legionella effector AnkX reveals the mechanism of phosphocholine transfer by the FIC domain
Abstract
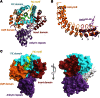
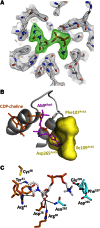
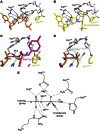
The FIC motif and the eukaryotic-like ankyrin repeats are found in many bacterial type IV effectors, yet little is known about how these domains enable bacteria to modulate host cell functions. Bacterial FIC domains typically bind ATP and transfer adenosine monophosphate moiety onto target proteins. The ankyrin repeat-containing protein AnkX encoded by the intracellular pathogen Legionella pneumophila is unique in that its FIC domain binds to CDP-choline and transfers a phosphocholine residue onto proteins in the Rab1 GTPase family. By determining the structures of unbound AnkX and AnkX with bound CDP-choline, CMP/phosphocholine and CMP, we demonstrate that the orientation of substrate binding in relation to the catalytic FIC motif enables this protein to function as a phosphocholinating enzyme rather than a nucleotidyl transferase. Additionally, the structure reveals that the ankyrin repeats mediate scaffolding interactions that resemble those found in protein-protein interactions, but are unprecedented in intramolecular interactions. Together with phosphocholination experiments, our structures unify a general phosphoryl transferase mechanism common to all FIC enzymes that should be conserved from bacteria to human.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures


References
-
- Adams PD, Afonine PV, Bunkoczi G, Chen VB, Davis IW, Echols N, Headd JJ, Hung LW, Kapral GJ, Grosse-Kunstleve RW, McCoy AJ, Moriarty NW, Oeffner R, Read RJ, Richardson DC, Richardson JS, Terwilliger TC, Zwart PH (2010) PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr D Biol Crystallogr 66: 213–221 - PMC - PubMed
-
- Blanc E, Roversi P, Vonrhein C, Flensburg C, Lea SM, Bricogne G (2004) Refinement of severely incomplete structures with maximum likelihood in BUSTER-TNT. Acta Crystallogr D Biol Crystallogr 60: 2210–2221 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

