Structure-function analysis of a broad specificity Populus trichocarpa endo-β-glucanase reveals an evolutionary link between bacterial licheninases and plant XTH gene products
- PMID: 23572521
- PMCID: PMC3668736
- DOI: 10.1074/jbc.M113.462887
Structure-function analysis of a broad specificity Populus trichocarpa endo-β-glucanase reveals an evolutionary link between bacterial licheninases and plant XTH gene products
Abstract
The large xyloglucan endotransglycosylase/hydrolase (XTH) gene family continues to be the focus of much attention in studies of plant cell wall morphogenesis due to the unique catalytic functions of the enzymes it encodes. The XTH gene products compose a subfamily of glycoside hydrolase family 16 (GH16), which also comprises a broad range of microbial endoglucanases and endogalactanases, as well as yeast cell wall chitin/β-glucan transglycosylases. Previous whole-family phylogenetic analyses have suggested that the closest relatives to the XTH gene products are the bacterial licheninases (EC 3.2.1.73), which specifically hydrolyze linear mixed linkage β(1→3)/β(1→4)-glucans. In addition to their specificity for the highly branched xyloglucan polysaccharide, XTH gene products are distinguished from the licheninases and other GH16 enzyme subfamilies by significant active site loop alterations and a large C-terminal extension. Given these differences, the molecular evolution of the XTH gene products in GH16 has remained enigmatic. Here, we present the biochemical and structural analysis of a unique, mixed function endoglucanase from black cottonwood (Populus trichocarpa), which reveals a small, newly recognized subfamily of GH16 members intermediate between the bacterial licheninases and plant XTH gene products. We postulate that this clade comprises an important link in the evolution of the large plant XTH gene families from a putative microbial ancestor. As such, this analysis provides new insights into the diversification of GH16 and further unites the apparently disparate members of this important family of proteins.
Keywords: Carbohydrate Processing; Cellulase; Enzyme Kinetics; Enzyme Structure; Glycoside Hydrolases; Licheninase; Plant Cell Wall; Polysaccharide; Xyloglucan; Xyloglucan Endotransglycosylase/Hydrolase (XTH).
Figures

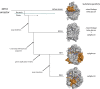
References
-
- Carpita N., McCann M. (2000) The cell wall. in Biochemistry and Molecular Biology of Plants (Buchanan B., Gruissem W., Jones R., eds) pp. 52–108, John Wiley & Sons, Inc., Somerset, NJ
-
- Himmel M. E., Ding S. Y., Johnson D. K., Adney W. S., Nimlos M. R., Brady J. W., Foust T. D. (2007) Biomass recalcitrance. Engineering plants and enzymes for biofuels production. Science 315, 804–807 - PubMed
-
- Ragauskas A. J., Williams C. K., Davison B. H., Britovsek G., Cairney J., Eckert C. A., Frederick W. J., Jr., Hallett J. P., Leak D. J., Liotta C. L., Mielenz J. R., Murphy R., Templer R., Tschaplinski T. (2006) The path forward for biofuels and biomaterials. Science 311, 484–489 - PubMed
-
- Cosgrove D. J. (2005) Growth of the plant cell wall. Nat. Rev. Mol. Cell Biol. 6, 850–861 - PubMed
-
- Vogel J. (2008) Unique aspects of the grass cell wall. Curr. Opin. Plant Biol. 11, 301–307 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

