MERISTEM DISORGANIZATION1 encodes TEN1, an essential telomere protein that modulates telomerase processivity in Arabidopsis
- PMID: 23572541
- PMCID: PMC3663272
- DOI: 10.1105/tpc.112.107425
MERISTEM DISORGANIZATION1 encodes TEN1, an essential telomere protein that modulates telomerase processivity in Arabidopsis
Abstract
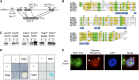
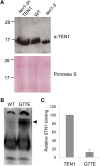
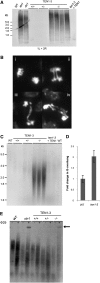
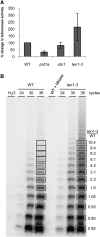
Telomeres protect chromosome ends from being recognized as DNA damage, and they facilitate the complete replication of linear chromosomes. CST [for CTC1(Cdc13)/STN1/TEN1] is a trimeric chromosome end binding complex implicated in both aspects of telomere function. Here, we characterize TEN1 in the flowering plant Arabidopsis thaliana. We report that TEN1 (for telomeric pathways in association with Stn1, which stands for suppressor of cdc thirteen) is encoded by a previously characterized gene, MERISTEM DISORGANIZATION1 (MDO1). A point mutation in MDO1, mdo1-1/ten1-3 (G77E), triggers stem cell differentiation and death as well as a constitutive DNA damage response. We provide biochemical and genetic evidence that ten1-3 is likely to be a null mutation. As with ctc1 and stn1 null mutants, telomere tracts in ten1-3 are shorter and more heterogeneous than the wild type. Mutants also exhibit frequent telomere fusions, increased single-strand telomeric DNA, and telomeric circles. However, unlike stn1 or ctc1 mutants, telomerase enzyme activity is elevated in ten1-3 mutants due to an increase in repeat addition processivity. In addition, TEN1 is detected at a significantly smaller fraction of telomeres than CTC1. These data indicate that TEN1 is critical for telomere stability and also plays an unexpected role in modulating telomerase enzyme activity.
Figures



Similar articles
-
Role of STN1 and DNA polymerase α in telomere stability and genome-wide replication in Arabidopsis.PLoS Genet. 2014 Oct 9;10(10):e1004682. doi: 10.1371/journal.pgen.1004682. eCollection 2014 Oct. PLoS Genet. 2014. PMID: 25299252 Free PMC article.
-
POT1a and components of CST engage telomerase and regulate its activity in Arabidopsis.PLoS Genet. 2014 Oct 16;10(10):e1004738. doi: 10.1371/journal.pgen.1004738. eCollection 2014 Oct. PLoS Genet. 2014. PMID: 25329641 Free PMC article.
-
STN1 protects chromosome ends in Arabidopsis thaliana.Proc Natl Acad Sci U S A. 2008 Dec 16;105(50):19815-20. doi: 10.1073/pnas.0807867105. Epub 2008 Dec 8. Proc Natl Acad Sci U S A. 2008. PMID: 19064932 Free PMC article.
-
Short telomeres: from dyskeratosis congenita to sporadic aplastic anemia and malignancy.Transl Res. 2013 Dec;162(6):353-63. doi: 10.1016/j.trsl.2013.05.003. Epub 2013 Jun 1. Transl Res. 2013. PMID: 23732052 Free PMC article. Review.
-
CST-Polα/Primase: the second telomere maintenance machine.Genes Dev. 2023 Jul 1;37(13-14):555-569. doi: 10.1101/gad.350479.123. Epub 2023 Jul 26. Genes Dev. 2023. PMID: 37495394 Free PMC article. Review.
Cited by
-
Progress in Human and Tetrahymena Telomerase Structure Determination.Annu Rev Biophys. 2017 May 22;46:199-225. doi: 10.1146/annurev-biophys-062215-011140. Epub 2017 Mar 15. Annu Rev Biophys. 2017. PMID: 28301767 Free PMC article. Review.
-
Genetic architecture of natural variation of telomere length in Arabidopsis thaliana.Genetics. 2015 Feb;199(2):625-35. doi: 10.1534/genetics.114.172163. Epub 2014 Dec 8. Genetics. 2015. PMID: 25488978 Free PMC article.
-
Extending the model of Arabidopsis telomere length and composition across Brassicaceae.Chromosome Res. 2014 Jun;22(2):153-66. doi: 10.1007/s10577-014-9423-y. Chromosome Res. 2014. PMID: 24846723
-
Role of STN1 and DNA polymerase α in telomere stability and genome-wide replication in Arabidopsis.PLoS Genet. 2014 Oct 9;10(10):e1004682. doi: 10.1371/journal.pgen.1004682. eCollection 2014 Oct. PLoS Genet. 2014. PMID: 25299252 Free PMC article.
-
Telomeres: an organized string linking plants and mammals.Biol Direct. 2024 Nov 20;19(1):119. doi: 10.1186/s13062-024-00558-y. Biol Direct. 2024. PMID: 39568075 Free PMC article. Review.
References
-
- Anderson B.H., et al. (2012). Mutations in CTC1, encoding conserved telomere maintenance component 1, cause Coats plus. Nat. Genet. 44: 338–342 - PubMed
-
- Armstrong S.J., Sanchez-Moran E., Franklin F.C. (2009). Cytological analysis of Arabidopsis thaliana meiotic chromosomes. Methods Mol. Biol. 558: 131–145 - PubMed
-
- Chen L.Y., Redon S., Lingner J. (2012). The human CST complex is a terminator of telomerase activity. Nature 488: 540–544 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

