Species identification and profiling of complex microbial communities using shotgun Illumina sequencing of 16S rRNA amplicon sequences
- PMID: 23579286
- PMCID: PMC3620293
- DOI: 10.1371/journal.pone.0060811
Species identification and profiling of complex microbial communities using shotgun Illumina sequencing of 16S rRNA amplicon sequences
Abstract
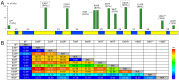
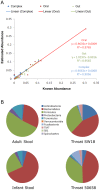
The high throughput and cost-effectiveness afforded by short-read sequencing technologies, in principle, enable researchers to perform 16S rRNA profiling of complex microbial communities at unprecedented depth and resolution. Existing Illumina sequencing protocols are, however, limited by the fraction of the 16S rRNA gene that is interrogated and therefore limit the resolution and quality of the profiling. To address this, we present the design of a novel protocol for shotgun Illumina sequencing of the bacterial 16S rRNA gene, optimized to amplify more than 90% of sequences in the Greengenes database and with the ability to distinguish nearly twice as many species-level OTUs compared to existing protocols. Using several in silico and experimental datasets, we demonstrate that despite the presence of multiple variable and conserved regions, the resulting shotgun sequences can be used to accurately quantify the constituents of complex microbial communities. The reconstruction of a significant fraction of the 16S rRNA gene also enabled high precision (>90%) in species-level identification thereby opening up potential application of this approach for clinical microbial characterization.
Conflict of interest statement
Figures
References
-
- Illumina Inc. (2008) Multiplexed Sequencing with the Illumina Genome Analyzer System. http://www.illumina.com/documents/products/datasheets/datasheet_sequenci....
-
- Baker GC, Smith JJ, Cowan DA (2003) Review and re-analysis of domain-specific 16S primers. J. Microbiol. Methods 55: 541–555. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

