3D phenotyping and quantitative trait locus mapping identify core regions of the rice genome controlling root architecture
- PMID: 23580618
- PMCID: PMC3645568
- DOI: 10.1073/pnas.1304354110
3D phenotyping and quantitative trait locus mapping identify core regions of the rice genome controlling root architecture
Abstract
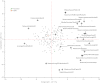
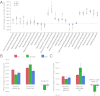
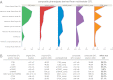
Identification of genes that control root system architecture in crop plants requires innovations that enable high-throughput and accurate measurements of root system architecture through time. We demonstrate the ability of a semiautomated 3D in vivo imaging and digital phenotyping pipeline to interrogate the quantitative genetic basis of root system growth in a rice biparental mapping population, Bala × Azucena. We phenotyped >1,400 3D root models and >57,000 2D images for a suite of 25 traits that quantified the distribution, shape, extent of exploration, and the intrinsic size of root networks at days 12, 14, and 16 of growth in a gellan gum medium. From these data we identified 89 quantitative trait loci, some of which correspond to those found previously in soil-grown plants, and provide evidence for genetic tradeoffs in root growth allocations, such as between the extent and thoroughness of exploration. We also developed a multivariate method for generating and mapping central root architecture phenotypes and used it to identify five major quantitative trait loci (r(2) = 24-37%), two of which were not identified by our univariate analysis. Our imaging and analytical platform provides a means to identify genes with high potential for improving root traits and agronomic qualities of crops.
Conflict of interest statement
The authors declare no conflict of interest.
Figures





References
-
- Munns R, et al. Wheat grain yield on saline soils is improved by an ancestral Na+ transporter gene. Nat Biotechnol. 2012;30:360–364. - PubMed
-
- Magalhaes JV, et al. A gene in the multidrug and toxic compound extrusion (MATE) family confers aluminum tolerance in sorghum. Nat Genet. 2007;39(9):1156–1161. - PubMed
-
- de Dorlodot S, et al. Root system architecture: Opportunities and constraints for genetic improvement of crops. Trends Plant Sci. 2007;12(10):474–481. - PubMed
-
- Ludlow MM, Muchow RC. 1990. Advances in Agronomy (Elsevier, Philadelphia)
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

