The cyclomodulin cycle inhibiting factor (CIF) alters cullin neddylation dynamics
- PMID: 23589306
- PMCID: PMC3663497
- DOI: 10.1074/jbc.M112.448258
The cyclomodulin cycle inhibiting factor (CIF) alters cullin neddylation dynamics
Abstract
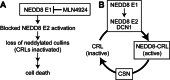
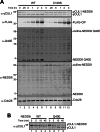
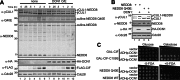
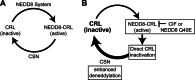
The bacterial effector protein cycle inhibiting factor (CIF) converts glutamine 40 of NEDD8 to glutamate (Q40E), causing cytopathic effects and inhibiting cell proliferation. Although these have been attributed to blocking the functions of cullin-RING ubiquitin ligases, how CIF modulates NEDD8-dependent signaling is unclear. Here we use conditional NEDD8-dependent yeast to explore the effects of CIF on cullin neddylation. Although CIF causes cullin deneddylation and the generation of free NEDD8 Q40E, inhibiting the COP9 signalosome (CSN) allows Q40E to form only on NEDD8 attached to cullins. In the presence of the CSN, NEDD8 Q40E is removed from cullins more rapidly than NEDD8, leading to a decrease in steady-state cullin neddylation. As NEDD8 Q40E is competent for cullin conjugation in the absence of functional CSN and with overexpression of the NEDD8 ligase Dcn1, our data are consistent with NEDD8 deamidation causing enhanced deneddylation of cullins by the CSN. This leads to a dramatic change in the extent of activated cullin-RING ubiquitin ligases.
Keywords: CIF; Cullin-RING Ubiquitin Ligases; NEDD8; Signal Transduction; Ubiquitin; Ubiquitin Ligase; Ubiquitin-like Proteins; Ubiquitination; Yeast.
Figures






Similar articles
-
Inhibition of cullin RING ligases by cycle inhibiting factor: evidence for interference with Nedd8-induced conformational control.J Mol Biol. 2011 Oct 21;413(2):430-7. doi: 10.1016/j.jmb.2011.08.030. Epub 2011 Aug 29. J Mol Biol. 2011. PMID: 21903097
-
The conserved protein DCN-1/Dcn1p is required for cullin neddylation in C. elegans and S. cerevisiae.Nature. 2005 Jun 30;435(7046):1257-61. doi: 10.1038/nature03662. Nature. 2005. PMID: 15988528
-
Neddylation and deneddylation regulate Cul1 and Cul3 protein accumulation.Nat Cell Biol. 2005 Oct;7(10):1014-20. doi: 10.1038/ncb1301. Epub 2005 Aug 28. Nat Cell Biol. 2005. PMID: 16127432
-
Protection of cullin-RING E3 ligases by CSN-UBP12.Trends Cell Biol. 2006 Jul;16(7):362-9. doi: 10.1016/j.tcb.2006.05.001. Epub 2006 Jun 9. Trends Cell Biol. 2006. PMID: 16762551 Review.
-
The COP9 signalosome and its role in plant development.Eur J Cell Biol. 2010 Feb-Mar;89(2-3):157-62. doi: 10.1016/j.ejcb.2009.11.021. Epub 2009 Dec 24. Eur J Cell Biol. 2010. PMID: 20036030 Review.
Cited by
-
Which bacterial toxins are worthy of validation as markers in colorectal cancer screening? A critical review.Biomed Pap Med Fac Univ Palacky Olomouc Czech Repub. 2022 Mar;166(1):1-11. doi: 10.5507/bp.2021.054. Epub 2021 Nov 5. Biomed Pap Med Fac Univ Palacky Olomouc Czech Repub. 2022. PMID: 34747413 Review.
-
Structural insights into the mechanism and inhibition of transglutaminase-induced ubiquitination by the Legionella effector MavC.Nat Commun. 2020 Apr 14;11(1):1774. doi: 10.1038/s41467-020-15645-7. Nat Commun. 2020. PMID: 32286321 Free PMC article.
-
Phylogenetic relationships of fluorescent pseudomonads deduced from the sequence analysis of 16S rRNA, Pseudomonas-specific and rpoD genes.3 Biotech. 2016 Jun;6(1):80. doi: 10.1007/s13205-016-0386-x. Epub 2016 Feb 23. 3 Biotech. 2016. PMID: 28330150 Free PMC article.
-
The Proteasome Lid Triggers COP9 Signalosome Activity during the Transition of Saccharomyces cerevisiae Cells into Quiescence.Biomolecules. 2019 Sep 4;9(9):449. doi: 10.3390/biom9090449. Biomolecules. 2019. PMID: 31487956 Free PMC article.
-
Exploitation of the host ubiquitin system by human bacterial pathogens.Nat Rev Microbiol. 2014 Jun;12(6):399-413. doi: 10.1038/nrmicro3259. Epub 2014 May 7. Nat Rev Microbiol. 2014. PMID: 24801936 Review.
References
-
- Cardozo T., Pagano M. (2004) The SCF ubiquitin ligase. Insights into a molecular machine. Nat. Rev. Mol. Cell Biol. 5, 739–751 - PubMed
-
- Deshaies R. J., Joazeiro C. A. (2009) RING domain E3 ubiquitin ligases. Annu. Rev. Biochem. 78, 399–434 - PubMed
-
- Zheng N., Schulman B. A., Song L., Miller J. J., Jeffrey P. D., Wang P., Chu C., Koepp D. M., Elledge S. J., Pagano M., Conaway R. C., Conaway J. W., Harper J. W., Pavletich N. P. (2002) Structure of the Cul1-Rbx1-Skp1-F boxSkp2 SCF ubiquitin ligase complex. Nature 416, 703–709 - PubMed
-
- Petroski M. D., Deshaies R. J. (2005) Function and regulation of cullin-RING ubiquitin ligases. Nat. Rev. Mol. Cell Biol. 6, 9–20 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

