Endoplasmic reticulum (ER) stress response and its physiological roles in plants
- PMID: 23591838
- PMCID: PMC3645738
- DOI: 10.3390/ijms14048188
Endoplasmic reticulum (ER) stress response and its physiological roles in plants
Abstract
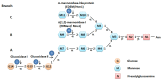
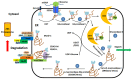
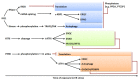
The endoplasmic reticulum (ER) stress response is a highly conserved mechanism that results from the accumulation of unfolded or misfolded proteins in the ER. The response plays an important role in allowing plants to sense and respond to adverse environmental conditions, such as heat stress, salt stress and pathogen infection. Since the ER is a well-controlled microenvironment for proper protein synthesis and folding, it is highly susceptible to stress conditions. Accumulation of unfolded or misfolded proteins activates a signaling pathway, called the unfolded protein response (UPR), which acts to relieve ER stress and, if unsuccessful, leads to cell death. Plants have two arms of the UPR signaling pathway, an arm involving the proteolytic processing of membrane-associated basic leucine zipper domain (bZIP) transcription factors and an arm involving RNA splicing factor, IRE1, and its mRNA target. These signaling pathways play an important role in determining the cell's fate in response to stress conditions.
Figures

Similar articles
-
Endoplasmic reticulum stress responses in plants.Annu Rev Plant Biol. 2013;64:477-99. doi: 10.1146/annurev-arplant-050312-120053. Epub 2013 Jan 7. Annu Rev Plant Biol. 2013. PMID: 23330794 Review.
-
Endoplasmic Reticulum Stress and Unfolded Protein Response Signaling in Plants.Int J Mol Sci. 2022 Jan 13;23(2):828. doi: 10.3390/ijms23020828. Int J Mol Sci. 2022. PMID: 35055014 Free PMC article. Review.
-
When is the unfolded protein response not the unfolded protein response?Plant Sci. 2017 Jul;260:139-143. doi: 10.1016/j.plantsci.2017.03.014. Epub 2017 Apr 5. Plant Sci. 2017. PMID: 28554471 Review.
-
Managing the protein folding demands in the endoplasmic reticulum of plants.New Phytol. 2016 Jul;211(2):418-28. doi: 10.1111/nph.13915. Epub 2016 Mar 14. New Phytol. 2016. PMID: 26990454 Review.
-
Misfolded proteins bind and activate death receptor 5 to trigger apoptosis during unresolved endoplasmic reticulum stress.Elife. 2020 Jan 6;9:e52291. doi: 10.7554/eLife.52291. Elife. 2020. PMID: 31904339 Free PMC article.
Cited by
-
Mitochondrial Defects Confer Tolerance against Cellulose Deficiency.Plant Cell. 2016 Sep;28(9):2276-2290. doi: 10.1105/tpc.16.00540. Epub 2016 Aug 19. Plant Cell. 2016. PMID: 27543091 Free PMC article.
-
Global Phosphoproteomic Analysis Reveals the Defense and Response Mechanisms of Jatropha Curcas Seedling under Chilling Stress.Int J Mol Sci. 2019 Jan 8;20(1):208. doi: 10.3390/ijms20010208. Int J Mol Sci. 2019. PMID: 30626061 Free PMC article.
-
Therapeutic Potential of Hydrogen Sulfide in Ischemia and Reperfusion Injury.Biomolecules. 2024 Jun 22;14(7):740. doi: 10.3390/biom14070740. Biomolecules. 2024. PMID: 39062455 Free PMC article. Review.
-
The THERMOSENSITIVE MALE STERILE 1 Interacts with the BiPs via DnaJ Domain and Stimulates Their ATPase Enzyme Activities in Arabidopsis.PLoS One. 2015 Jul 17;10(7):e0132500. doi: 10.1371/journal.pone.0132500. eCollection 2015. PLoS One. 2015. PMID: 26186593 Free PMC article.
-
StresSeed: The Unfolded Protein Response During Seed Development.Front Plant Sci. 2022 Mar 31;13:869008. doi: 10.3389/fpls.2022.869008. eCollection 2022. Front Plant Sci. 2022. PMID: 35432435 Free PMC article. Review.
References
-
- Lutcke H. Signal recognition particle (SRP), a ubiquitous initiator of protein translocation. Eur. J. Biochem. 1995;228:531–550. - PubMed
-
- Walter P., Lingappa V.R. Mechanism of protein translocation across the endoplasmic reticulum membrane. Annu. Rev. Cell Biol. 1986;2:499–516. - PubMed
-
- Walter P., Johnson A.E. Signal sequence recognition and protein targeting to the endoplasmic reticulum membrane. Annu. Rev. Cell Biol. 1994;10:87–119. - PubMed
-
- Marcinowski M., Holler M., Feige M.J., Baerend D., Lamb D.C., Buchner J. Substrate discrimination of the chaperone BiP by autonomous and cochaperone-regulated conformational transitions. Nat. Struct. Mol. Biol. 2011;18:150–158. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

