Programmed heterogeneity: epigenetic mechanisms in bacteria
- PMID: 23592777
- PMCID: PMC3656251
- DOI: 10.1074/jbc.R113.472274
Programmed heterogeneity: epigenetic mechanisms in bacteria
Abstract
Contrary to the traditional view that bacterial populations are clonal, single-cell analysis reveals that phenotypic heterogeneity is common in bacteria. Formation of distinct bacterial lineages appears to be frequent during adaptation to harsh environments, including the colonization of animals by bacterial pathogens. Formation of bacterial subpopulations is often controlled by epigenetic mechanisms that generate inheritable phenotypic diversity without altering the DNA sequence. Such mechanisms are diverse, ranging from relatively simple feedback loops to complex self-perpetuating DNA methylation patterns.
Keywords: Bacterial Genetics; DNA Methylation; DNA Methylation Pattern; DNA Methyltransferase; Epigenetics; Escherichia coli; Gene Regulation; Genetic Switch; Phase Variation; Reversible Bistability.
Figures
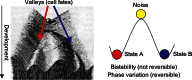


References
-
- Waddington C. H. (1957) The Strategy of the Genes, George Allen and Unwin, London
-
- Goldberg A. D., Allis C. D., Bernstein E. (2007) Epigenetics: a landscape takes shape. Cell 128, 635–638 - PubMed
-
- Errington J. (2003) Regulation of endospore formation in Bacillus subtilis. Nat. Rev. Microbiol. 1, 117–126 - PubMed
-
- Kirkpatrick C. L., Viollier P. H. (2012) Decoding Caulobacter development. FEMS Microbiol. Rev. 36, 193–205 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

