Evolution after introduction of a novel metabolic pathway consistently leads to restoration of wild-type physiology
- PMID: 23593025
- PMCID: PMC3616920
- DOI: 10.1371/journal.pgen.1003427
Evolution after introduction of a novel metabolic pathway consistently leads to restoration of wild-type physiology
Abstract
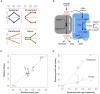
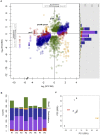
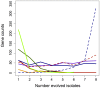
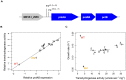
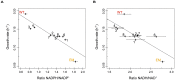
Organisms cope with physiological stressors through acclimatizing mechanisms in the short-term and adaptive mechanisms over evolutionary timescales. During adaptation to an environmental or genetic perturbation, beneficial mutations can generate numerous physiological changes: some will be novel with respect to prior physiological states, while others might either restore acclimatizing responses to a wild-type state, reinforce them further, or leave them unchanged. We examined the interplay of acclimatizing and adaptive responses at the level of global gene expression in Methylobacterium extorquens AM1 engineered with a novel central metabolism. Replacing central metabolism with a distinct, foreign pathway resulted in much slower growth than wild-type. After 600 generations of adaptation, however, eight replicate populations founded from this engineered ancestor had improved up to 2.5-fold. A comparison of global gene expression in wild-type, engineered, and all eight evolved strains revealed that the vast majority of changes during physiological adaptation effectively restored acclimatizing processes to wild-type expression states. On average, 93% of expression perturbations from the engineered strain were restored, with 70% of these occurring in perfect parallel across all eight replicate populations. Novel changes were common but typically restricted to one or a few lineages, and reinforcing changes were quite rare. Despite this, cases in which expression was novel or reinforced in parallel were enriched for loci harboring beneficial mutations. One case of parallel, reinforced changes was the pntAB transhydrogenase that uses NADH to reduce NADP(+) to NADPH. We show that PntAB activity was highly correlated with the restoration of NAD(H) and NADP(H) pools perturbed in the engineered strain to wild-type levels, and with improved growth. These results suggest that much of the evolved response to genetic perturbation was a consequence rather than a cause of adaptation and that physiology avoided "reinventing the wheel" by restoring acclimatizing processes to the pre-stressed state.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
Similar articles
-
Dissecting the genetic and metabolic mechanisms of adaptation to the knockout of a major metabolic enzyme in Escherichia coli.Proc Natl Acad Sci U S A. 2018 Jan 2;115(1):222-227. doi: 10.1073/pnas.1716056115. Epub 2017 Dec 18. Proc Natl Acad Sci U S A. 2018. PMID: 29255023 Free PMC article.
-
The soluble and membrane-bound transhydrogenases UdhA and PntAB have divergent functions in NADPH metabolism of Escherichia coli.J Biol Chem. 2004 Feb 20;279(8):6613-9. doi: 10.1074/jbc.M311657200. Epub 2003 Dec 3. J Biol Chem. 2004. PMID: 14660605
-
Distinct transcriptional regulation of the two Escherichia coli transhydrogenases PntAB and UdhA.Microbiology (Reading). 2016 Sep;162(9):1672-1679. doi: 10.1099/mic.0.000346. Epub 2016 Aug 2. Microbiology (Reading). 2016. PMID: 27488847
-
Evolution of global regulatory networks during a long-term experiment with Escherichia coli.Bioessays. 2007 Sep;29(9):846-60. doi: 10.1002/bies.20629. Bioessays. 2007. PMID: 17691099 Review.
-
Engineering of NADPH regenerators in Escherichia coli for enhanced biotransformation.Appl Microbiol Biotechnol. 2013 Apr;97(7):2761-72. doi: 10.1007/s00253-013-4750-z. Epub 2013 Feb 19. Appl Microbiol Biotechnol. 2013. PMID: 23420268 Review.
Cited by
-
Proteins Related to the Type I Secretion System Are Associated with Secondary SecA_DEAD Domain Proteins in Some Species of Planctomycetes, Verrucomicrobia, Proteobacteria, Nitrospirae and Chlorobi.PLoS One. 2015 Jun 1;10(6):e0129066. doi: 10.1371/journal.pone.0129066. eCollection 2015. PLoS One. 2015. PMID: 26030905 Free PMC article.
-
Experimental Design, Population Dynamics, and Diversity in Microbial Experimental Evolution.Microbiol Mol Biol Rev. 2018 Jul 25;82(3):e00008-18. doi: 10.1128/MMBR.00008-18. Print 2018 Sep. Microbiol Mol Biol Rev. 2018. PMID: 30045954 Free PMC article. Review.
-
The genomic landscape of compensatory evolution.PLoS Biol. 2014 Aug 26;12(8):e1001935. doi: 10.1371/journal.pbio.1001935. eCollection 2014 Aug. PLoS Biol. 2014. PMID: 25157590 Free PMC article.
-
Thermal Plasticity and Evolutionary Constraints in Bacillus: Implications for Climate Change Adaptation.Biology (Basel). 2024 Dec 23;13(12):1088. doi: 10.3390/biology13121088. Biology (Basel). 2024. PMID: 39765755 Free PMC article.
-
First-Step Mutations during Adaptation Restore the Expression of Hundreds of Genes.Mol Biol Evol. 2016 Jan;33(1):25-39. doi: 10.1093/molbev/msv228. Epub 2015 Oct 24. Mol Biol Evol. 2016. PMID: 26500250 Free PMC article.
References
-
- Cooper TF, Rozen DE, Lenski RE (2003) Parallel changes in gene expression after 20,000 generations of evolution in Escherichia coli . P Natl Acad Sci Usa 100: 1072–1077 doi:10.1073/pnas.0334340100. - DOI - PMC - PubMed
-
- Cooper TF, Remold SK, Lenski RE, Schneider D (2008) Expression profiles reveal parallel evolution of epistatic interactions involving the CRP regulon in Escherichia coli . PLoS Genet 4: e35 doi:10.1371/journal.pgen.0040035. - DOI - PMC - PubMed
-
- Gresham D, Desai MM, Tucker CM, Jenq HT, Pai DA, et al. (2008) The repertoire and dynamics of evolutionary adaptations to controlled nutrient-limited environments in yeast. PLoS Genet 4: e1000303 doi:10.1371/journal.pgen.1000303. - DOI - PMC - PubMed
-
- McDonald MJ, Gehrig SM, Meintjes PL, Zhang X-X, Rainey PB (2009) Adaptive divergence in experimental populations of Pseudomonas fluorescens. IV. Genetic constraints guide evolutionary trajectories in a parallel adaptive radiation. Genetics 183: 1041–1053 doi:10.1534/genetics.109.107110. - DOI - PMC - PubMed
-
- Kvitek DJ, Sherlock G (2011) Reciprocal sign epistasis between frequently experimentally evolved adaptive mutations causes a rugged fitness landscape. PLoS Genet 7: e1002056 doi:10.1371/journal.pgen.1002056. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

