Evolutionary paths of the cAMP-dependent protein kinase (PKA) catalytic subunits
- PMID: 23593352
- PMCID: PMC3625193
- DOI: 10.1371/journal.pone.0060935
Evolutionary paths of the cAMP-dependent protein kinase (PKA) catalytic subunits
Abstract
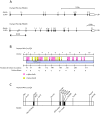
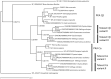
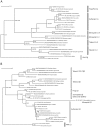
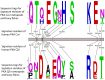
3',5'-cyclic adenosine monophosphate (cAMP) dependent protein kinase or protein kinase A (PKA) has served as a prototype for the large family of protein kinases that are crucially important for signal transduction in eukaryotic cells. The PKA catalytic subunits Cα and Cβ, encoded by the two genes PRKACA and PRKACB, respectively, are among the best understood and characterized human kinases. Here we have studied the evolution of this gene family in chordates, arthropods, mollusks and other animals employing probabilistic methods and show that Cα and Cβ arose by duplication of an ancestral PKA catalytic subunit in a common ancestor of vertebrates. The two genes have subsequently been duplicated in teleost fishes. The evolution of the PRKACG retroposon in simians was also investigated. Although the degree of sequence conservation in the PKA Cα/Cβ kinase family is exceptionally high, a small set of signature residues defining Cα and Cβ subfamilies were identified. These conserved residues might be important for functions that are unique to the Cα or Cβ clades. This study also provides a good example of a seemingly simple phylogenetic problem which, due to a very high degree of sequence conservation and corresponding weak phylogenetic signals, combined with problematic nonphylogenetic signals, is nontrivial for state-of-the-art probabilistic phylogenetic methods.
Conflict of interest statement
Figures
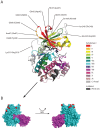
References
-
- Manning G, Whyte DB, Martinez R, Hunter T, Sudarsanam S (2002) The protein kinase complement of the human genome. Science 298: 1912–1934. - PubMed
-
- Skålhegg BS, Taskén K (2000) Specificity in the cAMP/PKA signaling pathway. Differential expression, regulation, and subcellular localization of subunits of PKA. Front Biosci 5: D678–D693. - PubMed
-
- Taylor SS, Yang J, Wu J, Haste NM, Radzio-Andzelm E, et al. (2004) PKA: a portrait of protein kinase dynamics. Biochim Biophys Acta 1697: 259–269. - PubMed
-
- Krebs EG, Beavo JA (1979) Phosphorylation-dephosphorylation of enzymes. Annu Rev Biochem 48: 923–959. - PubMed
-
- Kim C, Xuong NH, Taylor SS (2005) Crystal structure of a complex between the catalytic and regulatory (RIα) subunits of PKA. Science 307: 690–696. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

