Chromatin relaxation-mediated induction of p19INK4d increases the ability of cells to repair damaged DNA
- PMID: 23593412
- PMCID: PMC3625165
- DOI: 10.1371/journal.pone.0061143
Chromatin relaxation-mediated induction of p19INK4d increases the ability of cells to repair damaged DNA
Abstract
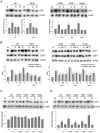
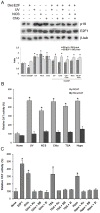
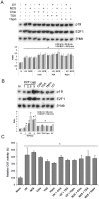
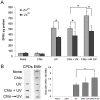
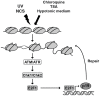
The maintenance of genomic integrity is of main importance to the survival and health of organisms which are continuously exposed to genotoxic stress. Cells respond to DNA damage by activating survival pathways consisting of cell cycle checkpoints and repair mechanisms. However, the signal that triggers the DNA damage response is not necessarily a direct detection of the primary DNA lesion. In fact, chromatin defects may serve as initiating signals to activate those mechanisms. If the modulation of chromatin structure could initiate a checkpoint response in a direct manner, this supposes the existence of specific chromatin sensors. p19INK4d, a member of the INK4 cell cycle inhibitors, plays a crucial role in regulating genomic stability and cell viability by enhancing DNA repair. Its expression is induced in cells injured by one of several genotoxic treatments like cis-platin, UV light or neocarzinostatin. Nevertheless, when exogenous DNA damaged molecules are introduced into the cell, this induction is not observed. Here, we show that p19INK4d is enhanced after chromatin relaxation even in the absence of DNA damage. This induction was shown to depend upon ATM/ATR, Chk1/Chk2 and E2F activity, as is the case of p19INK4d induction by endogenous DNA damage. Interestingly, p19INK4d improves DNA repair when the genotoxic damage is caused in a relaxed-chromatin context. These results suggest that changes in chromatin structure, and not DNA damage itself, is the actual trigger of p19INK4d induction. We propose that, in addition to its role as a cell cycle inhibitor, p19INK4d could participate in a signaling network directed to detecting and eventually responding to chromatin anomalies.
Conflict of interest statement
Figures


Similar articles
-
CDK2 and PKA mediated-sequential phosphorylation is critical for p19INK4d function in the DNA damage response.PLoS One. 2012;7(4):e35638. doi: 10.1371/journal.pone.0035638. Epub 2012 Apr 25. PLoS One. 2012. PMID: 22558186 Free PMC article.
-
E2F1 induces p19INK4d, a protein involved in the DNA damage response, following UV irradiation.Mol Cell Biochem. 2012 Jul;366(1-2):123-9. doi: 10.1007/s11010-012-1289-8. Epub 2012 Apr 3. Mol Cell Biochem. 2012. PMID: 22476863
-
ATM regulates ATR chromatin loading in response to DNA double-strand breaks.J Exp Med. 2006 Feb 20;203(2):297-303. doi: 10.1084/jem.20051923. Epub 2006 Feb 6. J Exp Med. 2006. PMID: 16461339 Free PMC article.
-
The ATM-Chk2 and ATR-Chk1 pathways in DNA damage signaling and cancer.Adv Cancer Res. 2010;108:73-112. doi: 10.1016/B978-0-12-380888-2.00003-0. Adv Cancer Res. 2010. PMID: 21034966 Review.
-
Protein kinases that regulate chromosome stability and their downstream targets.Genome Dyn. 2006;1:131-148. doi: 10.1159/000092505. Genome Dyn. 2006. PMID: 18724058 Review.
Cited by
-
Nuclear translocation of p19INK4d in response to oxidative DNA damage promotes chromatin relaxation.Mol Cell Biochem. 2015 Jan;398(1-2):63-72. doi: 10.1007/s11010-014-2205-1. Epub 2014 Sep 10. Mol Cell Biochem. 2015. PMID: 25204969
-
Retrieving the intracellular topology from multi-scale protein mobility mapping in living cells.Nat Commun. 2014 Jul 24;5:4494. doi: 10.1038/ncomms5494. Nat Commun. 2014. PMID: 25058002 Free PMC article.
-
Non-canonical functions of cell cycle cyclins and cyclin-dependent kinases.Nat Rev Mol Cell Biol. 2016 May;17(5):280-92. doi: 10.1038/nrm.2016.27. Epub 2016 Apr 1. Nat Rev Mol Cell Biol. 2016. PMID: 27033256 Free PMC article. Review.
-
To live or let die - complexity within the E2F1 pathway.Mol Cell Oncol. 2015 Jan 30;2(1):e970480. doi: 10.4161/23723548.2014.970480. eCollection 2015 Jan-Mar. Mol Cell Oncol. 2015. PMID: 27308406 Free PMC article. Review.
-
p19 INK4d controls hematopoietic stem cells in a cell-autonomous manner during genotoxic stress and through the microenvironment during aging.Stem Cell Reports. 2014 Dec 9;3(6):1085-102. doi: 10.1016/j.stemcr.2014.10.005. Epub 2014 Nov 20. Stem Cell Reports. 2014. PMID: 25458892 Free PMC article.
References
-
- Groth A, Rocha W, Verreault A, Almouzni G (2007) Chromatin challenges during DNA replication and repair. Cell 128: 721–733. - PubMed
-
- Loizou JI, Murr R, Finkbeiner MG, Sawan C, Wang ZQ, et al. (2006) Epigenetic information in chromatin: the code of entry for DNA repair. Cell Cycle 5: 696–701. - PubMed
-
- Celeste A, Fernandez-Capetillo O, Kruhlak MJ, Pilch DR, Staudt DW, et al. (2003) Histone H2AX phosphorylation is dispensable for the initial recognition of DNA breaks. Nat Cell Biol 5: 675–679. - PubMed
-
- Fernandez-Capetillo O, Celeste A, Nussenzweig A (2003) Focusing on foci: H2AX and the recruitment of DNA-damage response factors. Cell Cycle 2: 426–427. - PubMed
-
- Matsuoka S, Ballif BA, Smogorzewska A, McDonald ER, 3rd, Hurov KE, et al (2007) ATM and ATR substrate analysis reveals extensive protein networks responsive to DNA damage. Science 316: 1160–1166. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

