Sequence divergence in the Treponema denticola FhbB protein and its impact on factor H binding
- PMID: 23601078
- PMCID: PMC3785937
- DOI: 10.1111/omi.12027
Sequence divergence in the Treponema denticola FhbB protein and its impact on factor H binding
Abstract
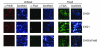
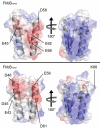
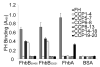
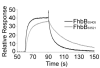
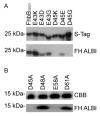
Treponema denticola is an anaerobic spirochete whose abundance in the subgingival crevice correlates with the development and severity of periodontal disease. The ability of T. denticola to survive and thrive in the hostile environment of the periodontal pocket is due, at least in part, to its ability to bind factor H (FH), a negative regulator of the alternative complement pathway. The FH binding protein of T. denticola has been identified as FhbB and its atomic structure has been determined. The interaction of FH with T. denticola is unique in that FH bound to the cell surface is cleaved by the T. denticola protease, dentilisin. It has been postulated that FH cleavage by T. denticola leads to immune dysregulation in periodontal pockets. In this study, we conduct a comparative assessment of the sequence, properties, structure and ligand binding kinetics of the FhbB proteins of strains 33521 and 35405. The biological outcome of the interaction of these strains with FH could differ significantly as 33521 lacks dentilisin activity. The data presented here offer insight into our understanding of the interactions of T. denticola with the host and its potential to influence disease progression.
Keywords: complement; factor H; immune evasion; periodontitis.
© 2013 John Wiley & Sons A/S. Published by John Wiley & Sons Ltd.
Figures
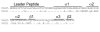
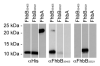

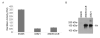
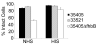
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

